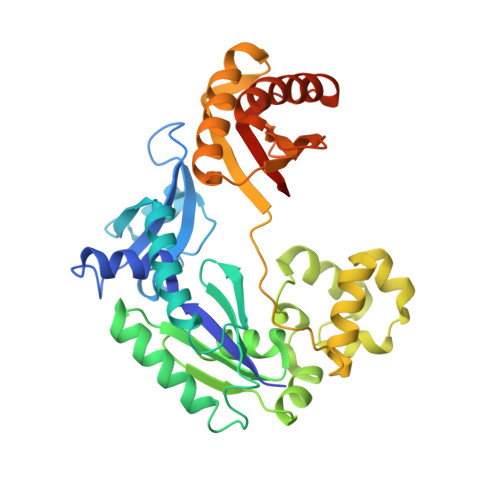
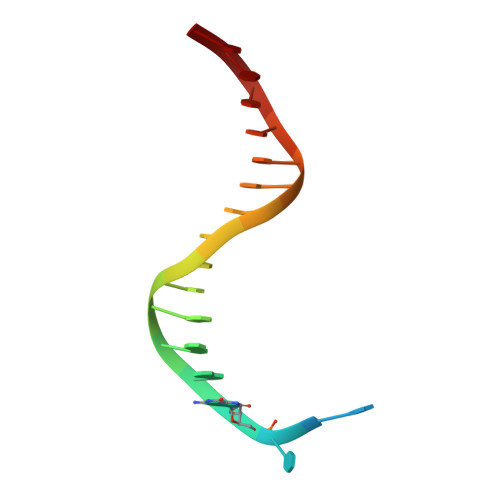
Efficient and High Fidelity Incorporation of Dctp Opposite 7,8-Dihydro-8-Oxodeoxyguanosine by Sulfolobus Solfataricus DNA Polymerase Dpo4
Zang, H., Irimia, A., Choi, J.-Y., Angel, K.C., Loukachevitch, L.V., Egli, M., Guengerich, F.P.(2006) J Biological Chem 281: 2358
- PubMed: 16306039 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M510889200
- Primary Citation Related Structures:
2C22, 2C28, 2C2D, 2C2E, 2C2R - PubMed Abstract:
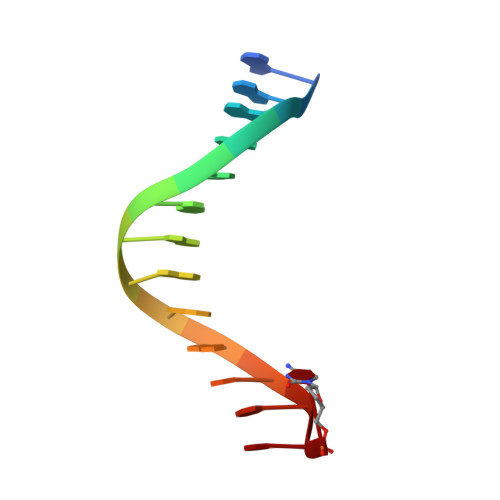
DNA polymerases insert dATP opposite the oxidative damage product 7,8-dihydro-8-oxodeoxyguanosine (8-oxoG) instead of dCTP, to the extent of >90% with some polymerases. Steady-state kinetics with the Y-family Sulfolobus solfataricus DNA polymerase IV (Dpo4) showed 90-fold higher incorporation efficiency of dCTP > dATP opposite 8-oxoG and 4-fold higher efficiency of extension beyond an 8-oxoG:C pair than an 8-oxoG:A pair. The catalytic efficiency for these events (with dCTP or C) was similar for G and 8-oxoG templates. Mass spectral analysis of extended DNA primers showed >/=95% incorporation of dCTP > dATP opposite 8-oxoG. Pre-steady-state kinetics showed faster rates of dCTP incorporation opposite 8-oxoG than G. The measured K(d)(,dCTP) was 15-fold lower for an oligonucleotide containing 8-oxoG than with G. Extension beyond an 8-oxoG:C pair was similar to G:C and faster than for an 8-oxoG:A pair, in contrast to other polymerases. The E(a) for dCTP insertion opposite 8-oxoG was lower than for opposite G. Crystal structures of Dpo4 complexes with oligonucleotides were solved with C, A, and G nucleoside triphosphates placed opposite 8-oxoG. With ddCTP, dCTP, and dATP the phosphodiester bonds were formed even in the presence of Ca(2+). The 8-oxoG:C pair showed classic Watson-Crick geometry; the 8-oxoG:A pair was in the syn:anti configuration, with the A hybridized in a Hoogsteen pair with 8-oxoG. With dGTP placed opposite 8-oxoG, pairing was not to the 8-oxoG but to the 5' C (and in classic Watson-Crick geometry), consistent with the low frequency of this frameshift event observed in the catalytic assays.
- Department of Biochemistry and Center in Molecular Toxicology, Vanderbilt University School of Medicine, 638 Robinson Research Building, 23rd and Pierce Avenues, Nashville, TN 37232-0146, USA.
Organizational Affiliation: