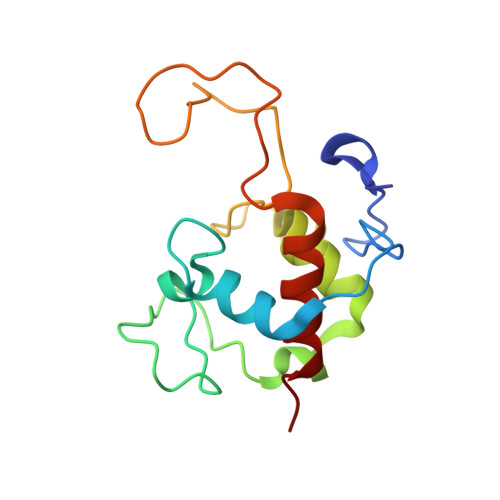
Structure of the Cytochrome Complex Soxxa of Paracoccus Pantotrophus, a Heme Enzyme Initiating Chemotrophic Sulfur Oxidation.
Dambe, T., Quentmeier, A., Rother, D., Friedrich, C., Scheidig, A.J.(2005) J Struct Biol 152: 229
- PubMed: 16297640 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2005.09.002
- Primary Citation Related Structures:
2C1D - PubMed Abstract:
The sulfur-oxidizing enzyme system (Sox) of the chemotroph Paracoccus pantotrophus is composed of several proteins, which together oxidize hydrogen sulfide, sulfur, thiosulfate or sulfite and transfers the gained electrons to the respiratory chain. The hetero-dimeric cytochrome c complex SoxXA functions as heme enzyme and links covalently the sulfur substrate to the thiol of the cysteine-138 residue of the SoxY protein of the SoxYZ complex. Here, we report the crystal structure of the c-type cytochrome complex SoxXA. The structure could be solved by molecular replacement and refined to a resolution of 1.9A identifying the axial heme-iron coordination involving an unusual Cys-251 thiolate of heme2. Distance measurements between the three heme groups provide deeper insight into the electron transport inside SoxXA and merge in a better understanding of the initial step of the aerobic sulfur oxidation process in chemotrophic bacteria.
- Max-Planck-Institut für Molekulare Physiologie, Abteilung Physikalische Biochemie, Otto-Hahn-Strasse 11, D-44225 Dortmund, Germany.
Organizational Affiliation: