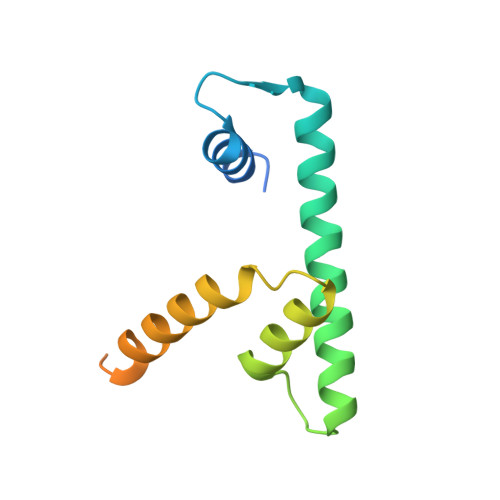
The Histone Fold Subunits of Drosophila Chrac Facilitate Nucleosome Sliding Through Dynamic DNA Interactions.
Hartlepp, K.F., Fernandez-Tornero, C., Eberharter, A., Grune, T., Muller, C.W., Becker, P.B.(2005) Mol Cell Biol 25: 9886
- PubMed: 16260604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.25.22.9886-9896.2005
- Primary Citation Related Structures:
2BYK, 2BYM - PubMed Abstract:
The chromatin accessibility complex (CHRAC) is an abundant, evolutionarily conserved nucleosome remodeling machinery able to catalyze histone octamer sliding on DNA. CHRAC differs from the related ACF complex by the presence of two subunits with molecular masses of 14 and 16 kDa, whose structure and function were not known. We determined the structure of Drosophila melanogaster CHRAC14-CHRAC16 by X-ray crystallography at 2.4-angstroms resolution and found that they dimerize via a variant histone fold in a typical handshake structure. In further analogy to histones, CHRAC14-16 contain unstructured N- and C-terminal tail domains that protrude from the handshake structure. A dimer of CHRAC14-16 can associate with the N terminus of ACF1, thereby completing CHRAC. Low-affinity interactions of CHRAC14-16 with DNA significantly improve the efficiency of nucleosome mobilization by limiting amounts of ACF. Deletion of the negatively charged C terminus of CHRAC16 enhances DNA binding 25-fold but leads to inhibition of nucleosome sliding, in striking analogy to the effect of the DNA chaperone HMGB1 on nucleosome sliding. The presence of a surface compatible with DNA interaction and the geometry of an H2A-H2B heterodimer may provide a transient acceptor site for DNA dislocated from the histone surface and therefore facilitate the nucleosome remodeling process.
- European Molecular Biology Laboratory, Grenoble Outstation, France.
Organizational Affiliation: