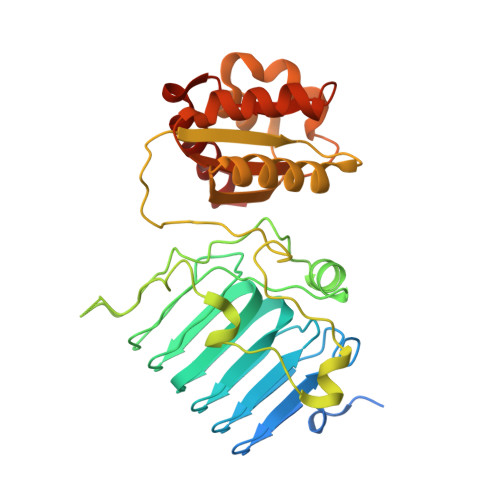
Crystal Structure of the Human Retinitis Pigmentosa 2 Protein and its Interaction with Arl3
Kuhnel, K., Veltel, S., Schlichting, I., Wittinghofer, A.(2006) Structure 14: 367
- PubMed: 16472755 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.11.008
- Primary Citation Related Structures:
2BX6 - PubMed Abstract:
The crystal structure of human retinitis pigmentosa 2 protein (RP2) was solved to 2.1 angstroms resolution. It consists of an N-terminal beta helix and a C-terminal ferredoxin-like alpha/beta domain. RP2 is functionally and structurally related to the tubulin-specific chaperone cofactor C. Seven of nine known RP2 missense mutations identified in patients are located in the beta helix domain, and most of them cluster to the hydrophobic core and are likely to destabilize the protein. Two residues, Glu138 and the catalytically important Arg118, are solvent-exposed and form a salt bridge, indicating that Glu138 might be critical for positioning Arg118 for catalysis. RP2 is a specific effector protein of Arl3. The N-terminal 34 residues and beta helix domain of RP2 are required for this interaction. The abilitities of RP2 to bind Arl3 and cause retinitis pigmentosa seem to be correlated, since both the R118H and E138G mutants show a drastically reduced affinity to Arl3.
- Max-Planck-Institut für Molekulare Physiologie, Abteilung Strukturelle Biologie, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany.
Organizational Affiliation: