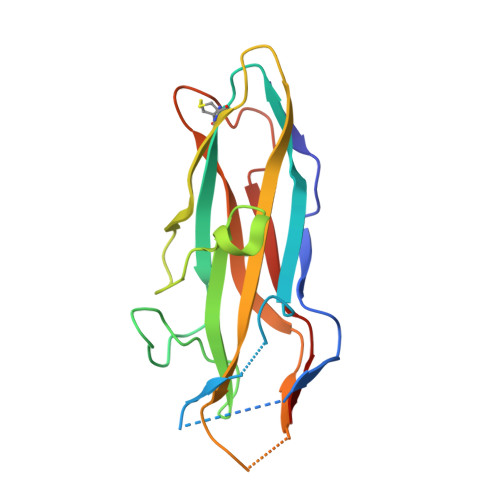
Impact of Natural Variation in Bacterial F17G Adhesins on Crystallization Behaviour.
Buts, L., Wellens, A., Van Molle, I., Wyns, L., Loris, R., Lahmann, M., Oscarson, S., De Greve, H., Bouckaert, J.(2005) Acta Crystallogr D Biol Crystallogr 61: 1149
- PubMed: 16041081 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905017038
- Primary Citation Related Structures:
1ZK5, 1ZPL, 2BS7, 2BS8, 2BSB, 2BSC - PubMed Abstract:
Since the introduction of structural genomics, the protein has been recognized as the most important variable in crystallization. Recent strategies to modify a protein to improve crystal quality have included rationally engineered point mutations, truncations, deletions and fusions. Five naturally occurring variants, differing in 1-18 amino acids, of the 177-residue lectin domain of the F17G fimbrial adhesin were expressed and purified in identical ways. For four out of the five variants crystals were obtained, mostly in non-isomorphous space groups, with diffraction limits ranging between 2.4 and 1.1 A resolution. A comparative analysis of the crystal-packing contacts revealed that the variable amino acids are often involved in lattice contacts and a single amino-acid substitution can suffice to radically change crystal packing. A statistical approach proved reliable to estimate the compatibilities of the variant sequences with the observed crystal forms. In conclusion, natural variation, universally present within prokaryotic species, is a valuable genetic resource that can be favourably employed to enhance the crystallization success rate with considerably less effort than other strategies.
- Laboratorium voor Ultrastructuur, Vlaams Interuniversitair Instituut voor Biotechnologie and Vrije Universiteit Brussel, Pleinlaan 2, B-1050 Brussel, Belgium.
Organizational Affiliation: