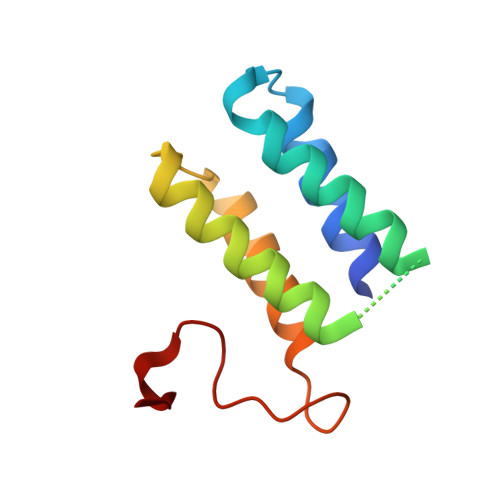
1.6-A Crystal Structure of Enta-Im: A Bacterial Immunity Protein Conferring Immunity to the Antimicrobial Activity of the Pediocin-Like Bacteriocin Enterocin A
Johnsen, L., Dalhus, B., Leiros, I., Nissen-Meyer, J.(2005) J Biological Chem 280: 19045
- PubMed: 15753083 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M501386200
- Primary Citation Related Structures:
2BL7, 2BL8 - PubMed Abstract:
Many Gram-positive bacteria produce ribosomally synthesized antimicrobial peptides, often termed bacteriocins. Genes encoding pediocin-like bacteriocins are generally cotranscribed with or in close vicinity to a gene encoding a cognate immunity protein that protects the bacteriocin-producer from their own bacteriocin. We present the first crystal structure of a pediocin-like immunity protein, EntA-im, conferring immunity to the bacteriocin enterocin A. Determination of the structure of this 103-amino acid protein revealed that it folds into an antiparallel four-helix bundle with a flexible C-terminal part. The fact that the immunity protein conferring immunity to carnobacteriocin B2 also consists of a four-helix bundle (Sprules, T., Kawulka, K. E., and Vederas, J. C. (2004) Biochemistry 43, 11740-11749) strongly indicates that this is a conserved structural motif in all pediocin-like immunity proteins. The C-terminal half of the immunity protein contains a region that recognizes the C-terminal half of the cognate bacteriocin, and the flexibility in the C-terminal end of the immunity protein might thus be an important characteristic that enables the immunity protein to interact with its cognate bacteriocin. By homology modeling of three other pediocin-like immunity proteins and calculation of the surface charge distribution for EntA-im and the three structure models, different charge distributions were observed. The differences in the latter part of helix 3, the beginning of helix 4, and the loop connecting these helices might also be of importance in determining the specificity.
- Program for Biochemistry and Molecular Biology, Department of Molecular Biosciences, University of Oslo, 0316 Oslo, Norway. line.johnsen@biokjemi.uio.no
Organizational Affiliation: