Structural Evidence for Non-Canonical Binding of Ca2+ to a Canonical EF-Hand of a Conventional Myosin.
Debreczeni, J.E., Farkas, L., Harmat, V., Hetenyi, C., Hajdu, I., Zavodszky, P., Kohama, K., Nyitray, L.(2005) J Biological Chem 280: 41458
- PubMed: 16227209 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M506315200
- Primary Citation Related Structures:
2BL0 - PubMed Abstract:
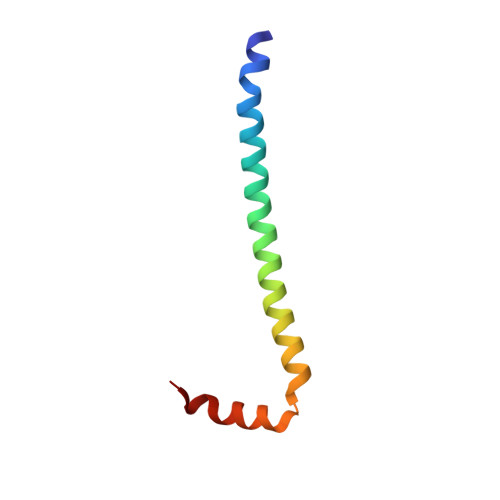
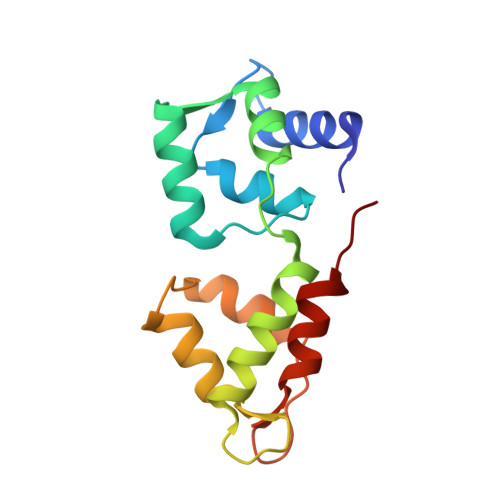
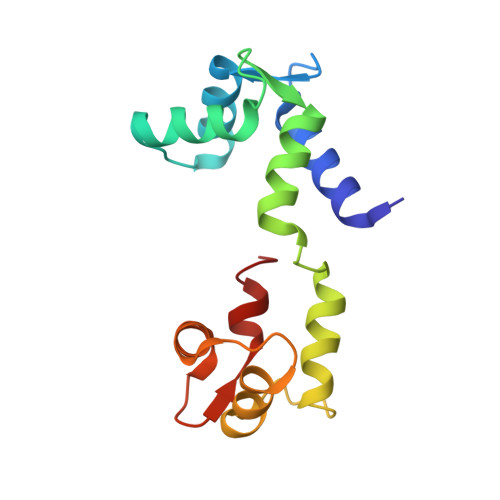
We have previously identified a single inhibitory Ca2+-binding site in the first EF-hand of the essential light chain of Physarum conventional myosin (Farkas, L., Malnasi-Csizmadia, A., Nakamura, A., Kohama, K., and Nyitray, L. (2003) J. Biol. Chem. 278, 27399-27405). As a general rule, conformation of the EF-hand-containing domains in the calmodulin family is "closed" in the absence and "open" in the presence of bound cations; a notable exception is the unusual Ca2+-bound closed domain in the essential light chain of the Ca2+-activated scallop muscle myosin. Here we have reported the 1.8 A resolution structure of the regulatory domain (RD) of Physarum myosin II in which Ca2+ is bound to a canonical EF-hand that is also in a closed state. The 12th position of the EF-hand loop, which normally provides a bidentate ligand for Ca2+ in the open state, is too far in the structure to participate in coordination of the ion. The structure includes a second Ca2+ that only mediates crystal contacts. To reveal the mechanism behind the regulatory effect of Ca2+, we compared conformational flexibilities of the liganded and unliganded RD. Our working hypothesis, i.e. the modulatory effect of Ca2+ on conformational flexibility of RD, is in line with the observed suppression of hydrogen-deuterium exchange rate in the Ca2+-bound form, as well as with results of molecular dynamics calculations. Based on this evidence, we concluded that Ca2+-induced change in structural dynamics of RD is a major factor in Ca2+-mediated regulation of Physarum myosin II activity.
- Department of Biochemistry, Eötvös Loránd University, Budapest H-1117, Hungary.
Organizational Affiliation: