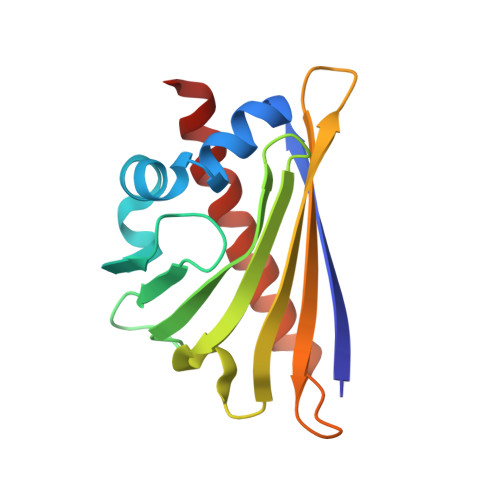
Crystal Structure of the Major Celery Allergen Api G 1: Molecular Analysis of Cross-Reactivity.
Schirmer, T., Hoffmann-Somergrube, K., Susani, M., Breiteneder, H., Markovic-Housley, Z.(2005) J Mol Biology 351: 1101
- PubMed: 16051263 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.06.054
- Primary Citation Related Structures:
2BK0 - PubMed Abstract:
Many patients who have been sensitised to pollen, display allergic symptoms after ingestion of certain plant food such as fresh fruit, vegetables and nuts. The cause is the cross-reactivity between structurally very similar major plant allergens. In particular, allergy to celery is very frequently associated with birch and mugwort pollen sensitization, known as to the birch-mugwort-celery syndrome. The crystal structure of the major celery allergen Api g 1, a homologue of the major birch pollen allergen Bet v 1, has been determined to a resolution of 2.9 A. The structure of Api g 1 is very similar to that of Bet v 1 with major differences occurring in the segment comprised of residues 23-45, preceding the well conserved glycine-rich P-loop, as well as in loops beta3-beta4 and beta5-beta6. In particular, Api g 1 lacks E45, which has been shown to be a crucial residue for antibody recognition in the crystal complex of Bet v 1 with the Fab fragment of a murine monoclonal IgG (BV16) antibody. The absence of E45 and the structural differences in the preceding segment suggest that this region of the Api g 1 surface is probably not responsible for the observed cross-reactivity with Bet v 1. A detailed analysis of the molecular surface in combination with sequence alignment revealed three conserved surface patches which may account for cross-reactivity with Bet v 1. Several residues of Bet v 1 which have been shown by mutagenesis studies to be involved in IgE recognition belong to these conserved surface regions. The structure of Api g 1 and the related epitope analysis provides a molecular basis for a better understanding of allergen cross-reactivity and may lead to the development of hypoallergens which would allow a safer immunotherapy.
- Department of Structural Biology, Biozentrum, University of Basel, CH-4056 Basel, Switzerland.
Organizational Affiliation: