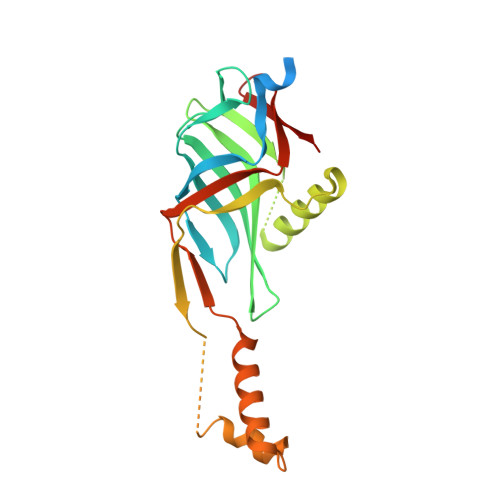
Structures of Two Core Subunits of the Bacterial Type Iv Secretion System, Virb8 from Brucella Suis and Comb10 from Helicobacter Pylori
Terradot, L., Bayliss, R., Oomen, C., Leonard, G., Baron, C., Waksman, G.(2005) Proc Natl Acad Sci U S A 102: 4596
- PubMed: 15764702 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0408927102
- Primary Citation Related Structures:
2BHM, 2BHV - PubMed Abstract:
Type IV secretion systems (T4SSs) are commonly used secretion machineries in Gram-negative bacteria. They are used in the infection of human, animal, or plant cells and the propagation of antibiotic resistance. The T4SS apparatus spans both membranes of the bacterium and generally is composed of 12 proteins, named VirB1-11 and VirD4 after proteins of the canonical Agrobacterium tumefaciens T4SS. The periplasmic core complex of VirB8/VirB10 structurally and functionally links the cytoplasmic NTPases of the system with its outer membrane and pilus components. Here we present crystal structures of VirB8 of Brucella suis, the causative agent of brucellosis, and ComB10, a VirB10 homolog of Helicobacter pylori, the causative agent of gastric ulcers. The structures of VirB8 and ComB10 resemble known folds, albeit with novel secondary-structure modifications unique to and conserved within their respective families. Both proteins crystallized as dimers, providing detailed predictions about their self associations. These structures make a substantial contribution to the repertoire of T4SS component structures and will serve as springboards for future functional and protein-protein interaction studies by using knowledge-based site-directed and deletion mutagenesis.
- Institute of Structural Molecular Biology, Malet Street, London WC1E 7HX, UK.
Organizational Affiliation: