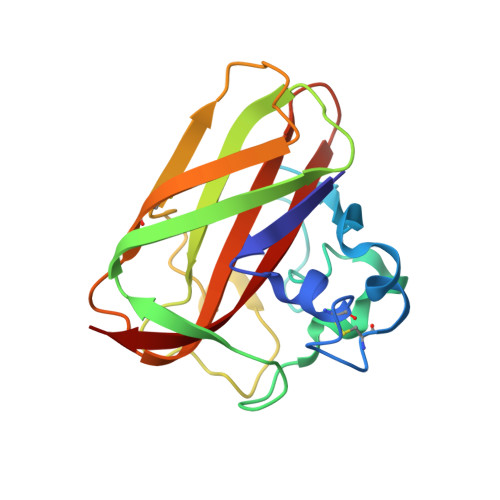
Crystal Structure and Binding Properties of the Serratia Marcescens Chitin-Binding Protein Cbp21
Vaaje-Kolstad, G., Houston, D.R., Eijsink, V.G.H., Van Aalten, D.M.F.(2005) J Biological Chem 280: 11313
- PubMed: 15590674
- DOI: https://doi.org/10.1074/jbc.M407175200
- Primary Citation Related Structures:
2BEM, 2BEN - PubMed Abstract:
Chitin proteins are commonly found in bacteria that utilize chitin as a source of energy. CBP21 is a chitin-binding protein from Serratia marcescens, a Gram-negative soil bacterium capable of efficient chitin degradation. When grown on chitin, S. marcescens secretes large amounts of CBP21, along with chitin-degrading enzymes. In an attempt to understand the molecular mechanism of CBP21 action, we have determined its crystal structure at 1.55 angstroms resolution. This is the first structure to be solved of a family 33 carbohydrate-binding module. The structure reveals a "budded" fibronectin type III fold consisting of two beta-sheets, arranged as a beta-sheet sandwich, with a 65-residue "bud" consisting of three short helices, located between beta-strands 1 and 2. Remarkably, conserved aromatic residues that have been suggested previously to play a role in chitin binding were mainly found in the interior of the protein, seemingly incapable of interacting with chitin, whereas the structure revealed a surface patch of highly conserved, mainly hydrophilic residues. The roles of six of these conserved surface-exposed residues (Tyr-54, Glu-55, Glu-60, His-114, Asp-182, and Asn-185) were probed by site-directed mutagenesis and subsequent binding studies. All single point mutations lowered the affinity of CBP21 for beta-chitin, as shown by 3-8-fold increases in the apparent binding constant. Thus, binding of CBP21 to chitin seems to be mediated primarily by conserved, solvent-exposed, polar side chains.
- Department of Chemistry, Biotechnology, and Food Science, Postbox 5003, Agricultural University of Norway, N-1432 As, Norway.
Organizational Affiliation: