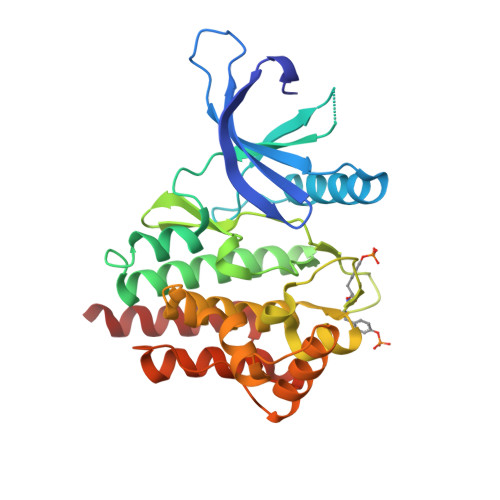
The structural basis of Janus kinase 2 inhibition by a potent and specific pan-Janus kinase inhibitor.
Lucet, I.S., Fantino, E., Styles, M., Bamert, R., Patel, O., Broughton, S.E., Walter, M., Burns, C.J., Treutlein, H., Wilks, A.F., Rossjohn, J.(2006) Blood 107: 176-183
- PubMed: 16174768
- DOI: https://doi.org/10.1182/blood-2005-06-2413
- Primary Citation of Related Structures:
2B7A - PubMed Abstract:
JAK2, a member of the Janus kinase (JAK) family of protein tyrosine kinases (PTKs), is an important intracellular mediator of cytokine signaling. Mutations of the JAK2 gene are associated with hematologic cancers, and aberrant JAK activity is also associated with a number of immune diseases, including rheumatoid arthritis. Accordingly, the development of JAK2-specific inhibitors has tremendous clinical relevance. Critical to the function of JAK2 is its PTK domain. We report the 2.0 A crystal structure of the active conformation of the JAK2 PTK domain in complex with a high-affinity, pan-JAK inhibitor that appears to bind via an induced fit mechanism. This inhibitor, the tetracyclic pyridone 2-tert-butyl-9-fluoro-3,6-dihydro-7H-benz[h]-imidaz[4,5-f]isoquinoline-7-1, was buried deep within a constricted ATP-binding site, in which extensive interactions, including residues that are unique to JAK2 and the JAK family, are made with the inhibitor. We present a structural basis of high-affinity JAK-specific inhibition that will undoubtedly provide an invaluable tool for the further design of novel, potent, and specific therapeutics against the JAK family.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria 3800, Australia.
Organizational Affiliation: