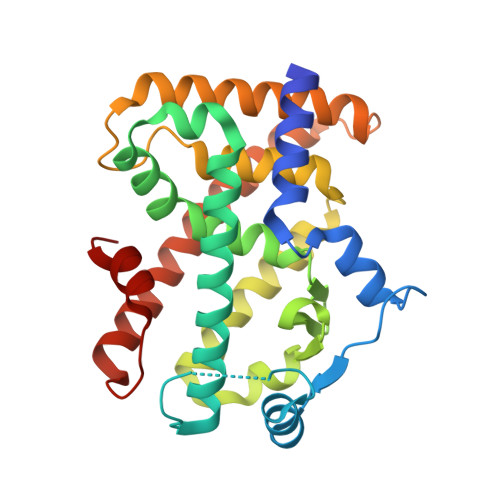
Recombinant Human PPAR-beta/delta Ligand-binding Domain is Locked in an Activated Conformation by Endogenous Fatty Acids
Fyffe, S.A., Alphey, M.S., Buetow, L., Smith, T.K., Ferguson, M.A.J., Sorensen, M.D., Bjorkling, F., Hunter, W.N.(2006) J Mol Biology 356: 1005-1013
- PubMed: 16405912 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.12.047
- Primary Citation Related Structures:
2AWH, 2B50 - PubMed Abstract:
High-resolution crystallographic structures of recombinant human peroxisome proliferator-activated receptor ligand-binding domain (isotype beta/delta) reveal a fatty acid in the binding site. Mass spectrometry confirmed the presence of C16:0, C16:1, C18:0 and C18:1 in a ratio of approximately 3:2:1:4 with 11, Z-octadecenoic acid (cis-vaccenic acid) identified as the predominant species. These are endogenous fatty acids acquired from the bacterial expression system, and serve to lock the ligand-binding domain into the activated conformation. A requirement for crystal growth, the additive n-heptyl-beta-d-glucopyranoside, binds near the activation function helix where recognition of co-activator proteins occurs. Our observations suggest potential physiological ligands for human PPAR-beta/delta and highlight that reported binding studies must be treated with caution unless endogenous fatty acids have been removed from the sample prior to analysis.
- Division of Biological Chemistry and Molecular Microbiology, School of Life Sciences, University of Dundee, Dundee DD1 5EH, UK.
Organizational Affiliation: