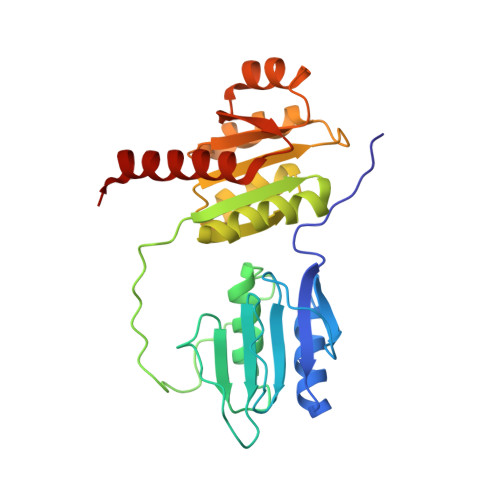
A New Structural Domain in the Escherichia coli RcsC Hybrid Sensor Kinase Connects Histidine Kinase and Phosphoreceiver Domains
Rogov, V.V., Rogova, N.Y., Bernhard, F., Koglin, A., Lohr, F., Dotsch, V.(2006) J Mol Biology 364: 68-79
- PubMed: 17005198 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2006.07.052
- Primary Citation Related Structures:
2AYX, 2AYY, 2AYZ - PubMed Abstract:
The Rcs signalling pathway controls a variety of physiological functions like capsule synthesis, cell division or motility in prokaryotes. The Rcs regulation cascade, involving a multi-step phosphorelay between the two membrane-bound hybrid sensor kinases RcsC and RcsD and the global regulator RcsB, is, up to now, one of the most complicated regulatory systems in bacteria. To understand the structural basis of Rcs signal transduction, NMR spectroscopy was employed to determine the solution structure of the RcsC C terminus, possessing a phosphoreceiver domain (RcsC-PR), and a region previously described as a long linker between the histidine kinase domain of RcsC (RcsC-HK) and the RcsC-PR. We have found that the linker region comprises an independent structural domain of a new alpha/beta organization, which we named RcsC-ABL domain (Alpha/Beta/Loop). The ABL domain appears to be a conserved and unique structural element of RcsC-like kinases with no significant sequence homology to other proteins. The second domain of the C terminus, the RcsC-PR domain, represents a well-folded CheY-like phosphoreceiver domain with the central parallel beta-sheet covered with two alpha-helical layers on both sides. We have mapped the interaction of RcsC-ABL and RcsC-PR with the histidine phosphotransfer domain (HPt) of RcsD. In addition we have characterized the interaction with and the conformational effects of Mg2+ and the phosphorylation mimetic BeF(-)(3) on RcsC-ABL and RcsC-PR.
- Institute of Biophysical Chemistry, Centre for Biomolecular Magnetic Resonance, JW Goethe-University, Frankfurt-am-Main, Germany.
Organizational Affiliation: