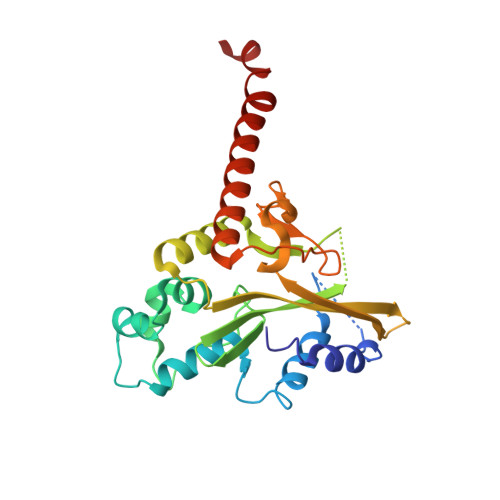
DNA-induced conformational changes in Type II restriction endonucleases: The structure of unliganded HincII
Horton, N.C., Little, E.J.(2005) J Mol Biology 351: 76-88
- PubMed: 15993893
- DOI: https://doi.org/10.1016/j.jmb.2005.05.063
- Primary Citation Related Structures:
2AUD - PubMed Abstract:
The 2.1A crystal structure of the unliganded type II restriction endonuclease, HincII, is described. Although the asymmetric unit contains only a single monomer, crystal lattice contacts bring two monomers together to form a dimer very similar to that found in the DNA bound form. Comparison with the published DNA bound structure reveals that the DNA binding pocket is expanded in the unliganded structure. Comparison of the unliganded and DNA liganded structures reveals a simple rotation of subunits by 11 degrees each, or 22 degrees total, to a more closed structure around the bound DNA. Comparison of this conformational change to that observed in the other type II restriction endonucleases where DNA bound and unliganded forms have both been characterized, shows considerable variation in the types of conformational changes that can occur. The conformational changes in three can be described by a simple rotation of subunits, while in two others both rotation and translation of subunits relative to one another occurs. In addition, the endonucleases having subunits that undergo the greatest amount of rotation upon DNA binding are found to be those that distort the bound DNA the least, suggesting that DNA bending may be less facile in dimers possessing greater flexibility.
- Department of Biochemistry and Molecular Biophysics, University of Arizona, Tucson, AZ 85721, USA.
Organizational Affiliation: