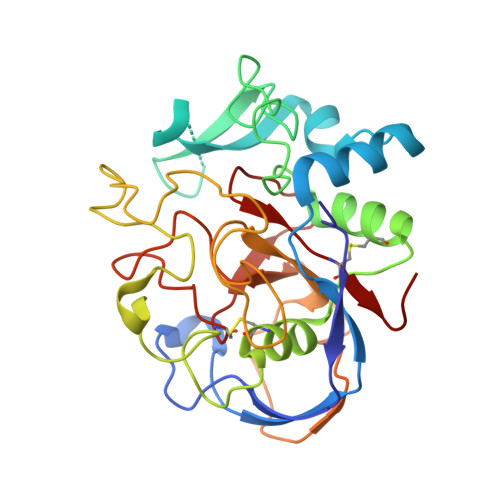
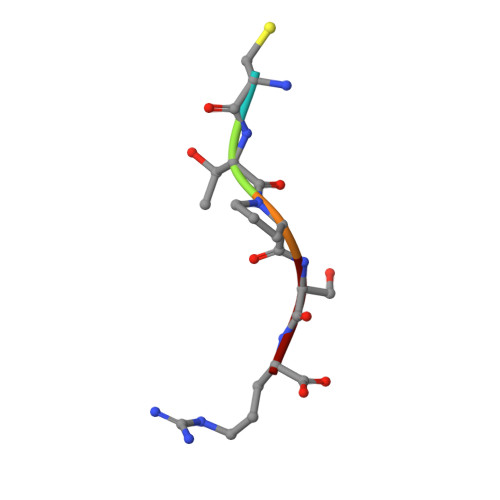
A general binding mechanism for all human sulfatases by the formylglycine-generating enzyme
Roeser, D., Preusser-Kunze, A., Schmidt, B., Gasow, K., Wittmann, J.G., Dierks, T., von Figura, K., Rudolph, M.G.(2006) Proc Natl Acad Sci U S A 103: 81-86
- PubMed: 16368756 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0507592102
- Primary Citation Related Structures:
2AFT, 2AFY, 2AII, 2AIJ, 2AIK - PubMed Abstract:
The formylglycine (FGly)-generating enzyme (FGE) uses molecular oxygen to oxidize a conserved cysteine residue in all eukaryotic sulfatases to the catalytically active FGly. Sulfatases degrade and remodel sulfate esters, and inactivity of FGE results in multiple sulfatase deficiency, a fatal disease. The previously determined FGE crystal structure revealed two crucial cysteine residues in the active site, one of which was thought to be implicated in substrate binding. The other cysteine residue partakes in a novel oxygenase mechanism that does not rely on any cofactors. Here, we present crystal structures of the individual FGE cysteine mutants and employ chemical probing of wild-type FGE, which defined the cysteines to differ strongly in their reactivity. This striking difference in reactivity is explained by the distinct roles of these cysteine residues in the catalytic mechanism. Hitherto, an enzyme-substrate complex as an essential cornerstone for the structural evaluation of the FGly formation mechanism has remained elusive. We also present two FGE-substrate complexes with pentamer and heptamer peptides that mimic sulfatases. The peptides isolate a small cavity that is a likely binding site for molecular oxygen and could host reactive oxygen intermediates during cysteine oxidation. Importantly, these FGE-peptide complexes directly unveil the molecular bases of FGE substrate binding and specificity. Because of the conserved nature of FGE sequences in other organisms, this binding mechanism is of general validity. Furthermore, several disease-causing mutations in both FGE and sulfatases are explained by this binding mechanism.
- Department of Molecular Structural Biology, University of Göttingen, Justus-von-Liebig-Weg 11, D-37077 Göttingen, Germany.
Organizational Affiliation: