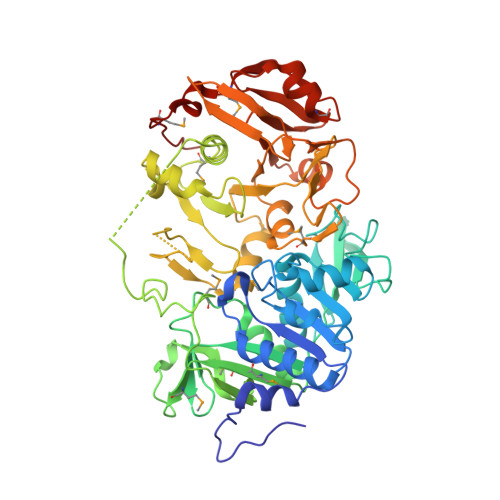
Crystallographic trapping of the glutamyl-CoA thioester intermediate of family I CoA transferases.
Rangarajan, E.S., Li, Y., Ajamian, E., Iannuzzi, P., Kernaghan, S.D., Fraser, M.E., Cygler, M., Matte, A.(2005) J Biological Chem 280: 42919-42928
- PubMed: 16253988 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M510522200
- Primary Citation Related Structures:
2AHU, 2AHV, 2AHW - PubMed Abstract:
Coenzyme A transferases are involved in a broad range of biochemical processes in both prokaryotes and eukaryotes, and exhibit a diverse range of substrate specificities. The YdiF protein from Escherichia coli O157:H7 is an acyl-CoA transferase of unknown physiological function, and belongs to a large sequence family of CoA transferases, present in bacteria to humans, which utilize oxoacids as acceptors. In vitro measurements showed that YdiF displays enzymatic activity with short-chain acyl-CoAs. The crystal structures of YdiF and its complex with CoA, the first co-crystal structure for any Family I CoA transferase, have been determined and refined at 1.9 and 2.0 A resolution, respectively. YdiF is organized into tetramers, with each monomer having an open alpha/beta structure characteristic of Family I CoA transferases. Co-crystallization of YdiF with a variety of CoA thioesters in the absence of acceptor carboxylic acid resulted in trapping a covalent gamma-glutamyl-CoA thioester intermediate. The CoA binds within a well defined pocket at the N- and C-terminal domain interface, but makes contact only with the C-terminal domain. The structure of the YdiF complex provides a basis for understanding the different catalytic steps in the reaction of Family I CoA transferases.
- Department of Biochemistry, McGill University, Quebec, Canada.
Organizational Affiliation: