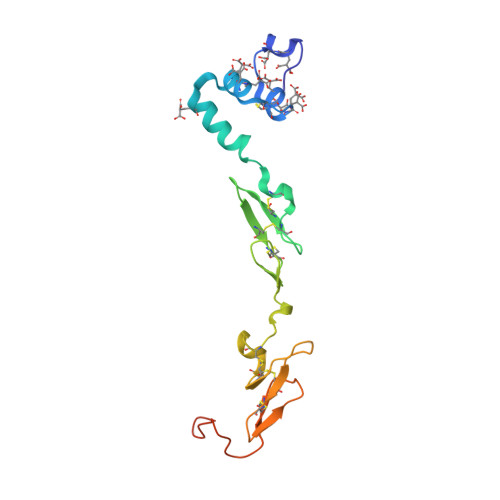
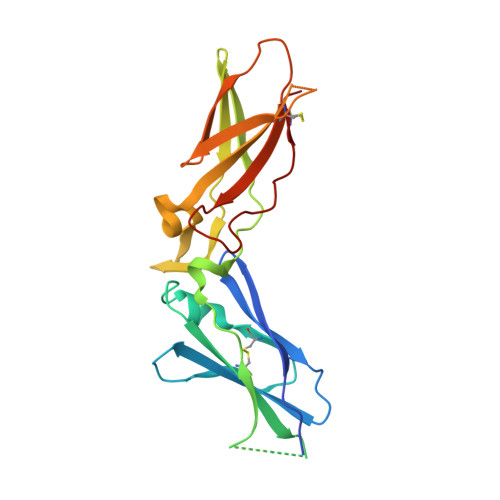
The discovery of fluoropyridine-based inhibitors of the Factor VIIa/TF complex.
Kohrt, J.T., Filipski, K.J., Cody, W.L., Cai, C., Dudley, D.A., Van Huis, C.A., Willardsen, J.A., Rapundalo, S.T., Saiya-Cork, K., Leadley, R.J., Narasimhan, L., Zhang, E., Whitlow, M., Adler, M., McLean, K., Chou, Y.L., McKnight, C., Arnaiz, D.O., Shaw, K.J., Light, D.R., Edmunds, J.J.(2005) Bioorg Med Chem Lett 15: 4752-4756
- PubMed: 16125385 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2005.07.059
- Primary Citation Related Structures:
2AEI - PubMed Abstract:
The activated Factor VII/tissue factor complex (FVIIa/TF) plays a key role in the formation of blood clots. Inhibition of this complex may lead to new antithrombotic drugs. An X-ray crystal structure of a fluoropyridine-based FVIIa/TF inhibitor bound in the active site of the enzyme complex suggested that incorporation of substitution at the 5-position of the hydroxybenzoic acid side chain could lead to the formation of more potent inhibitors through interactions with the S1'/S2' pocket.
- Pfizer Global Research and Development, Michigan Laboratories, 2800 Plymouth Road, Ann Arbor, MI 48105, USA. jeffrey.kohrt@pfizer.com
Organizational Affiliation: