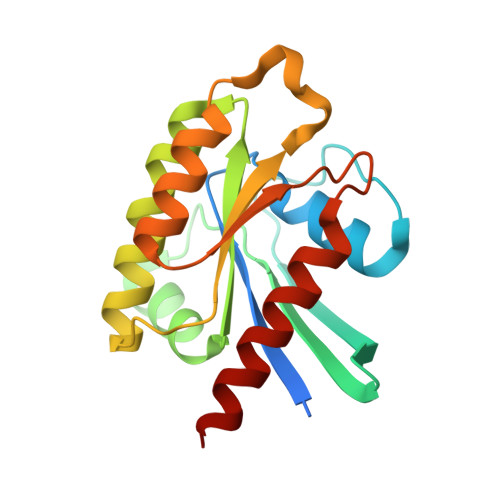
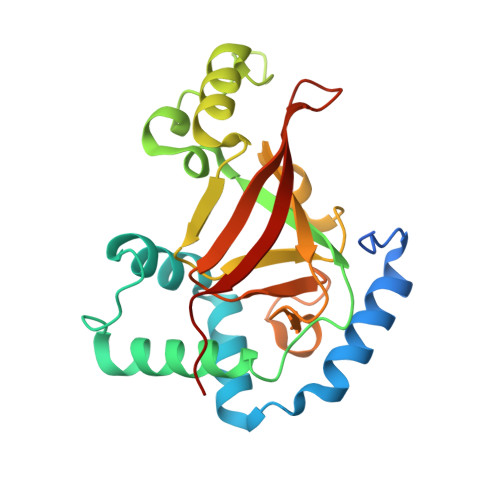
Crystal structure of the C3bot-RalA complex reveals a novel type of action of a bacterial exoenzyme.
Pautsch, A., Vogelsgesang, M., Trankle, J., Herrmann, C., Aktories, K.(2005) EMBO J 24: 3670-3680
- PubMed: 16177825 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600813
- Primary Citation Related Structures:
2A78, 2A9K - PubMed Abstract:
C3 exoenzymes from bacterial pathogens ADP-ribosylate and inactivate low-molecular-mass GTPases of the Rho subfamily. Ral, a Ras subfamily GTPase, binds the C3 exoenzymes from Clostridium botulinum and C. limosum with high affinity without being a substrate for ADP ribosylation. In the complex, the ADP-ribosyltransferase activity of C3 is blocked, while binding of NAD and NAD-glycohydrolase activity remain. Here we report the crystal structure of C3 from C. botulinum in a complex with GDP-bound RalA at 1.8 A resolution. C3 binds RalA with a helix-loop-helix motif that is adjacent to the active site. A quaternary complex with NAD suggests a mode for ADP-ribosyltransferase inhibition. Interaction of C3 with RalA occurs at a unique interface formed by the switch-II region, helix alpha3 and the P loop of the GTPase. C3-binding stabilizes the GDP-bound conformation of RalA and blocks nucleotide release. Our data indicate that C. botulinum exoenzyme C3 is a single-domain toxin with bifunctional properties targeting Rho GTPases by ADP ribosylation and Ral by a guanine nucleotide dissociation inhibitor-like effect, which blocks nucleotide exchange.
- Structural Research, Department of Integrated Lead Discovery, Boehringer Ingelheim Pharma GmbH & Co. KG, Biberach an der Riss, Germany.
Organizational Affiliation: