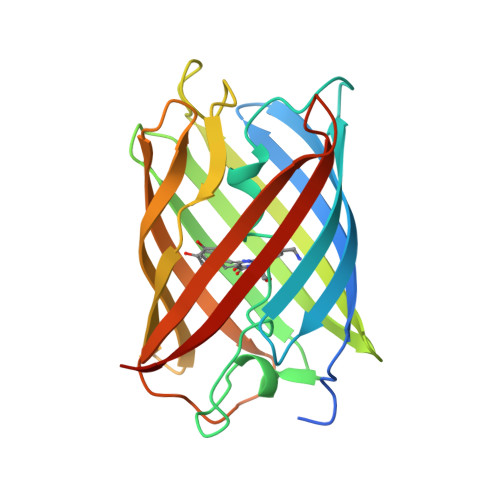
Crystal structures and mutational analysis of amFP486, a cyan fluorescent protein from Anemonia majano
Henderson, J.N., Remington, S.J.(2005) Proc Natl Acad Sci U S A 102: 12712-12717
- PubMed: 16120682 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0502250102
- Primary Citation Related Structures:
2A46, 2A47, 2A48 - PubMed Abstract:
Fluorescent proteins isolated from coral reef organisms can be roughly grouped into four color classes by emission, cyan, green, yellow, and red. To gain insight into the structural basis for cyan emission, the crystal structure of amFP486 (lambda(em)max = 486 nm) was determined by molecular replacement, and the model was refined at 1.65-A resolution. The electron density map reveals a chromophore formed from the tripeptide sequence -K-Y-G- that is indistinguishable from that of GFP (lambda(em)max = 509 nm). However, the chromophore environment closely parallels those of the yellow- and red-shifted homologs zFP538, DsRed, and eqFP611. Mutagenesis was performed for Glu-150, Ala-165, His-199, and Glu-217, which are immediately adjacent to the chromophore. His-199 and Ala-165 are key side chains responsible for the blue shift, presumably by localizing chromophore charge density on the phenolate moiety. Furthermore, in the H199T mutant the fluorescence quantum yield is reduced by a factor of approximately 110. The crystal structures of H199T (lambda(em)max = 515 nm) and E150Q (lambda(em)max = 506 nm) were determined. Remarkably, the H199T structure reveals that the stacking interaction of His-199 with the chromophore also controls the fluorescence efficiency, because the chromophore is statistically distributed in a 1:1 ratio between cis (fluorescent) and trans (nonfluorescent) conformations.
- Department of Physics and Institute of Molecular Biology, University of Oregon, Eugene, OR 97403, USA.
Organizational Affiliation: