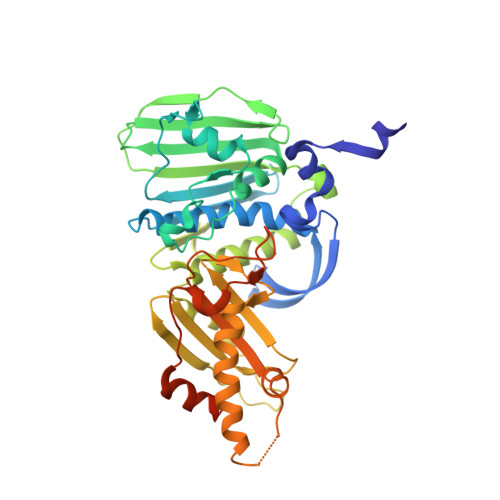
Nucleotide-dependent Domain Movement in the ATPase Domain of a Human Type IIA DNA Topoisomerase.
Wei, H., Ruthenburg, A.J., Bechis, S.K., Verdine, G.L.(2005) J Biological Chem 280: 37041-37047
- PubMed: 16100112 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M506520200
- Primary Citation Related Structures:
1ZXM, 1ZXN - PubMed Abstract:
Type IIA DNA topoisomerases play multiple essential roles in the management of higher-order DNA structure, including modulation of topological state, chromosome segregation, and chromatin condensation. These diverse physiologic functions are all accomplished through a common molecular mechanism, wherein the protein catalyzes transient cleavage of a DNA duplex (the G-segment) to yield a double-stranded gap through which another duplex (the T-segment) is passed. The overall process is orchestrated by the opening and closing of molecular "gates" in the topoisomerase structure, which is regulated by ATP binding, hydrolysis, and release of ADP and inorganic phosphate. Here we present two crystal structures of the ATPase domain of human DNA topoisomerase IIalpha in different nucleotide-bound states. Comparison of these structures revealed rigid-body movement of the structural modules within the ATPase domain, suggestive of the motions of a molecular gate.
- Department of Chemistry and Chemical Biology, Harvard University, Cambridge, Massachusetts 02138, USA.
Organizational Affiliation: