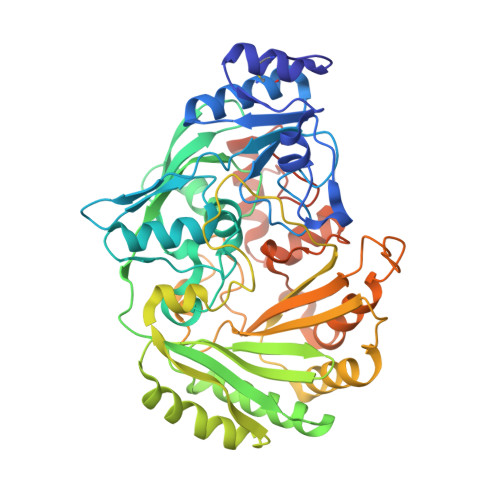
Crystal structure of glucooligosaccharide oxidase from Acremonium strictum: a novel flavinylation of 6-S-cysteinyl, 8alpha-N1-histidyl FAD
Huang, C.H., Lai, W.L., Lee, M.H., Chen, C.J., Vasella, A., Tsai, Y.C., Liaw, S.H.(2005) J Biological Chem 280: 38831-38838
- PubMed: 16154992
- DOI: https://doi.org/10.1074/jbc.M506078200
- Primary Citation of Related Structures:
1ZR6, 2AXR - PubMed Abstract:
Glucooligosaccharide oxidase from Acremonium strictum has been screened for potential applications in oligosaccharide acid production and alternative carbohydrate detection, because it catalyzes the oxidation of glucose, maltose, lactose, cellobiose and cello- and maltooligosaccharides. We report the crystal structures of the enzyme and of its complex with an inhibitor, 5-amino-5-deoxy- cellobiono-1,5-lactam at 1.55- and 1.98-A resolution, respectively. Unexpectedly, the protein structure demonstrates the first known double attachment flavinylation, 6-S-cysteinyl, 8alpha-N1-histidyl FAD. The FAD cofactor is cross-linked to the enzyme via the C(6) atom and the 8alpha-methyl group of the isoalloxazine ring with Cys(130) and His(70), respectively. This sugar oxidase possesses an open carbohydrate-binding groove, allowing the accommodation of higher oligosaccharides. The complex structure suggests that this enzyme may prefer a beta-d-glucosyl residue at the reducing end with the conserved Tyr(429) acting as a general base to abstract the OH(1) proton in concert with the H(1) hydride transfer to the flavin N(5). Finally, a detailed comparison illustrates the structural conservation as well as the divergence between this protein and its related flavoenzymes.
- Structural Biology Program, Institute of Biochemistry, and Faculty of Life Science, National Yang-Ming University, Taipei 11221, Taiwan.
Organizational Affiliation: