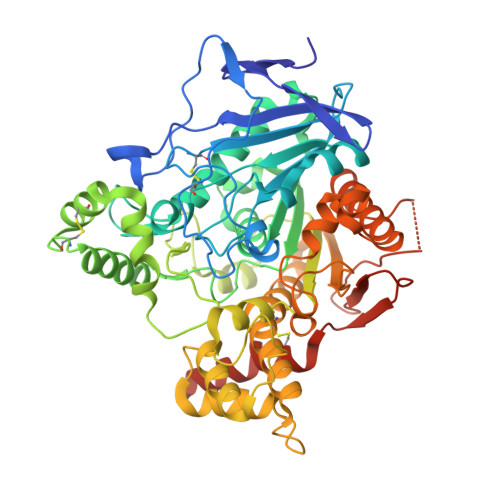
Crystal Packing Mediates Enantioselective Ligand Recognition at the Peripheral Site of Acetylcholinesterase
Haviv, H., Wong, D.M., Greenblatt, H.M., Carlier, P.R., Pang, Y.P., Silman, I., Sussman, J.L.(2005) J Am Chem Soc 127: 11029-11036
- PubMed: 16076210 Search on PubMed
- DOI: https://doi.org/10.1021/ja051765f
- Primary Citation Related Structures:
1ZGB, 1ZGC - PubMed Abstract:
Recently, alkylene-linked heterodimers of tacrine (1) and 5-amino-5,6,7,8-tetrahydroquinolinone (2, hupyridone) were shown to exhibit higher acetylcholinesterase (AChE) inhibition than either monomeric 1 or 2. Such inhibitors are potential drug candidates for ameliorating the cognitive decrements in early Alzheimer patients. In an attempt to understand the inhibition mechanism of one such dimer, (RS)-(+/-)-N-9-(1,2,3,4-tetrahydroacridinyl)-N'-5-[5,6,7,8-tetrahydro-2'(1'H)-quinolinonyl]-1,10-diaminodecane [(RS)-(+/-)-3] bisoxalate, the racemate was soaked in trigonal Torpedo californica AChE (TcAChE) crystals, and the X-ray structure of the resulting complex was solved to 2.30 A resolution. Its structure revealed the 1 unit bound to the "anionic" subsite of the active site, near the bottom of the active-site gorge, as seen for the 1/TcAChE complex. Interestingly, only the (R)-enantiomer of the 2 unit was seen in the peripheral "anionic" site (PAS) at the top of the gorge, and was hydrogen-bonded to the side chains of residues belonging to an adjacent, symmetry-related AChE molecule covering the gorge entrance. When the same racemate was soaked in orthorhombic crystals of TcAChE, in which the entrance to the gorge is more exposed, the crystal structure of the corresponding complex revealed no substantial enantiomeric selectivity. This observation suggests that the apparent enantiomeric selectivity of trigonal crystals of TcAChE for (R)-3 is mainly due to crystal packing, resulting in preferential binding of one enantiomeric inhibitor both to its "host" enzyme and to its neighbor in the asymmetric unit, rather than to steric constraints imposed by the geometry of the active-site gorge.
- Departments of Structural Biology and Neurobiology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: