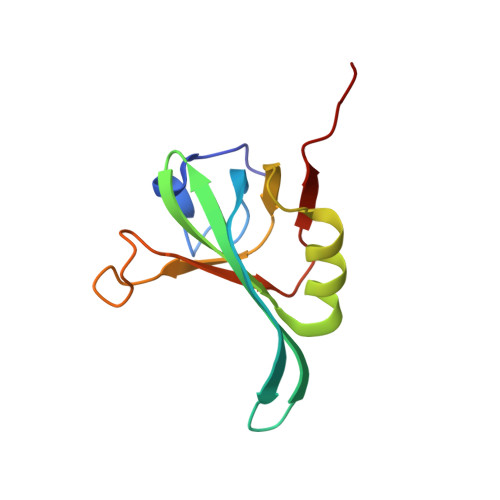
Solution structure of the DNA-binding domain of RPA from Saccharomyces cerevisiae and its interaction with single-stranded DNA and SV40 T antigen
Park, C.J., Lee, J.H., Choi, B.S.(2005) Nucleic Acids Res 33: 4172-4181
- PubMed: 16043636 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gki736
- Primary Citation Related Structures:
1YNX - PubMed Abstract:
Replication protein A (RPA) is a three-subunit complex with multiple roles in DNA metabolism. DNA-binding domain A in the large subunit of human RPA (hRPA70A) binds to single-stranded DNA (ssDNA) and is responsible for the species-specific RPA-T antigen (T-ag) interaction required for Simian virus 40 replication. Although Saccharomyces cerevisiae RPA70A (scRPA70A) shares high sequence homology with hRPA70A, the two are not functionally equivalent. To elucidate the similarities and differences between these two homologous proteins, we determined the solution structure of scRPA70A, which closely resembled the structure of hRPA70A. The structure of ssDNA-bound scRPA70A, as simulated by residual dipolar coupling-based homology modeling, suggested that the positioning of the ssDNA is the same for scRPA70A and hRPA70A, although the conformational changes that occur in the two proteins upon ssDNA binding are not identical. NMR titrations of hRPA70A with T-ag showed that the T-ag binding surface is separate from the ssDNA-binding region and is more neutral than the corresponding part of scRPA70A. These differences might account for the species-specific nature of the hRPA70A-T-ag interaction. Our results provide insight into how these two homologous RPA proteins can exhibit functional differences, but still both retain their ability to bind ssDNA.
- Department of Chemistry, National Creative Research Initiative Center, Korea Advanced Institute of Science and Technology 373-1, Guseong-dong, Yuseong-gu, Daejon 305-701, Korea.
Organizational Affiliation: