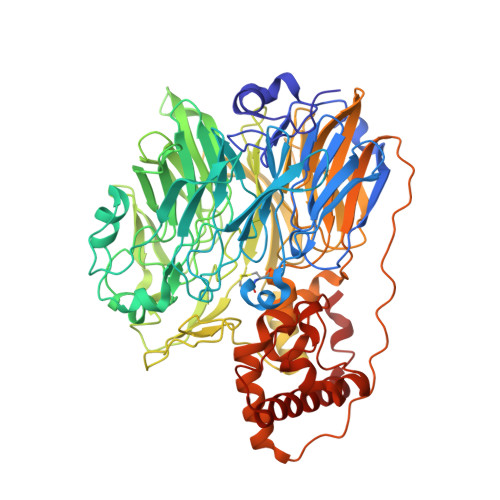
Molecular cloning and structural analysis of quinohemoprotein alcohol dehydrogenase ADH-IIG from Pseudomonas putida HK5
Toyama, H., Chen, Z.W., Fukumoto, M., Adachi, O., Matsushita, K., Mathews, F.S.(2005) J Mol Biology 352: 91-104
- PubMed: 16061256
- DOI: https://doi.org/10.1016/j.jmb.2005.06.078
- Primary Citation Related Structures:
1YIQ - PubMed Abstract:
Depending on the alcohols used as growth substrates, Pseudomonas putida HK5 produces two distinct quinohemoprotein alcohol dehydrogenases, ADH-IIB and ADH-IIG, both of which contain pyrroloquinoline quinone (PQQ) and heme c as the prosthetic groups but show different substrate specificities, especially for diol substrates. Molecular cloning of the gene of ADH-IIB and its crystal structure are already reported. Here, molecular cloning of the gene, qgdA, and solution of the three-dimensional structure of ADH-IIG are reported. The enzyme consists of 718 amino acid residues including a signal sequence of 29 amino acid residues. The PQQ domain is highly homologous to other quinoproteins, especially to quinohemoproteins. The crystal structure of ADH-IIG, determined at 2.2A resolution, shows that the overall structure and the amino acid residues involved in PQQ binding are quite similar to ADH-IIB and to another quinohemoprotein ADH, qhEDH from Comamonas testosteroni. However, the lengths of the linker regions connecting the PQQ and the cytochrome domains are different from each other, leading to a significant difference in orientation of the cytochrome domain with respect to the PQQ domain. Apart from ADH-IIB and qhEDH, ADH-IIG has an extra 12-residue helix within loop 3 in the PQQ domain and an extra 3(10) helix in the C terminus of the cytochrome domain, and both helices appear parallel and linked by a hydrogen bond. The amino acid residues contacting substrate/product in the crystal structures are also different among them. In the crystal structure of ADH-IIG with 1,2-propanediol, one of the hydroxyl groups of the substrate forms a hydrogen bond with O5 of PQQ and OD1 of Asp300, and the other interacts with a water molecule and with NE2 of Trp386, the corresponding residue of which is not found in ADH-IIB and qhEDH, and might be the residue responsible for making ADH-IIG prefer diol substrates.
- Department of Biological Chemistry, Faculty of Agriculture, Yamaguchi University, Yamaguchi 753-8515, Japan. hirot@yamaguchi-u.ac.jp
Organizational Affiliation: