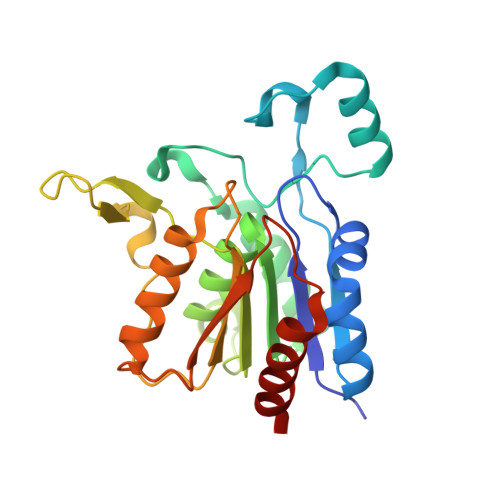
Crystal structure of yeast YHR049W/FSH1, a member of the serine hydrolase family.
Quevillon-Cheruel, S., Leulliot, N., Graille, M., Hervouet, N., Coste, F., Zelwer, C., Janin, J., Van Tilbeurgh, H.(2005) Protein Sci 14: 1350-1356
- PubMed: 15802654
- DOI: https://doi.org/10.1110/ps.051415905
- Primary Citation Related Structures:
1YCD - PubMed Abstract:
Yhr049w/FSH1 was recently identified in a combined computational and experimental proteomics analysis for the detection of active serine hydrolases in yeast. This analysis suggested that FSH1 might be a serine-type hydrolase belonging to the broad functional alphabeta-hydrolase superfamily. In order to get insight into the molecular function of this gene, it was targeted in our yeast structural genomics project. The crystal structure of the protein confirms that it contains a Ser/His/Asp catalytic triad that is part of a minimal alpha/beta-hydrolase fold. The architecture of the putative active site and analogies with other protein structures suggest that FSH1 may be an esterase. This finding was further strengthened by the unexpected presence of a compound covalently bound to the catalytic serine in the active site. Apparently, the enzyme was trapped with a reactive compound during the purification process.
- Institut de Biochimie et de Biophysique Moléculaire et Cellulaire (CNRS-UMR 8619), Université Paris-Sud, Bâtiment 430, 91405 Orsay, France.
Organizational Affiliation: