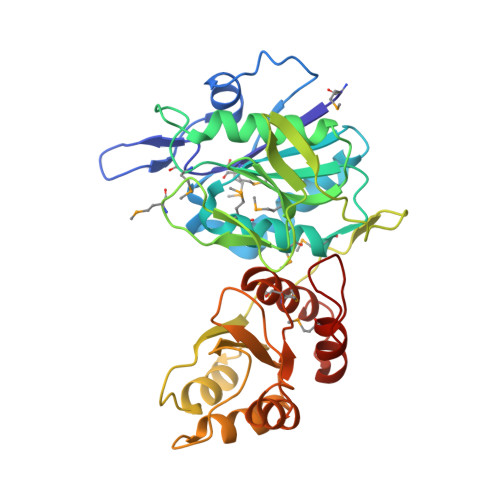
Structure of the type I L-asparaginase from the hyperthermophilic archaeon Pyrococcus horikoshii at 2.16 angstroms resolution.
Yao, M., Yasutake, Y., Morita, H., Tanaka, I.(2005) Acta Crystallogr D Biol Crystallogr 61: 294-301
- PubMed: 15735339
- DOI: https://doi.org/10.1107/S0907444904032950
- Primary Citation of Related Structures:
1WLS - PubMed Abstract:
The crystal structure of the L-asparaginase from the hyperthermophilic archaeon Pyrococcus horikoshii (PhA) was determined by the multiwavelength anomalous diffraction (MAD) method and was refined to a resolution of 2.16 angstroms with a crystallographic R factor and free R factor of 21.1 and 25.3%, respectively. This is the first report of the three-dimensional structure of a type I L-asparaginase. These enyzmes are known as cytosolic L-asparaginases with lower affinities for substrate than the type II L-asparaginases. Although the overall fold of PhA was closely related to the structure of the well characterized type II L-asparaginase, structural differences were also detected. PhA forms a homodimer that corresponds to half the homotetramer of type II L-asparaginases. Structure comparison at the active site reveals that most catalytic residues are conserved except for two residues that recognize the amino group of the substrate. Additionally, a remarkable structural difference is found in the so-called 'active-site flexible loop'. In PhA this loop is stabilized by beta-hairpin formation and by elaborate interactions with the type-I-specific alpha-helical region derived from the other subunit forming the PhA dimer. The flexible loop of the type II enzyme is considered to serve as a mobile gate to the active site. Therefore, the loop stabilization observed in the PhA structure may cause limitation of the access of the substrate to the active site.
- Division of Biological Sciences, Graduate School of Science, Hokkaido University, Kita-10, Nishi-8, Kita-ku, Sapporo, Hokkaido 060-0810, Japan.
Organizational Affiliation: