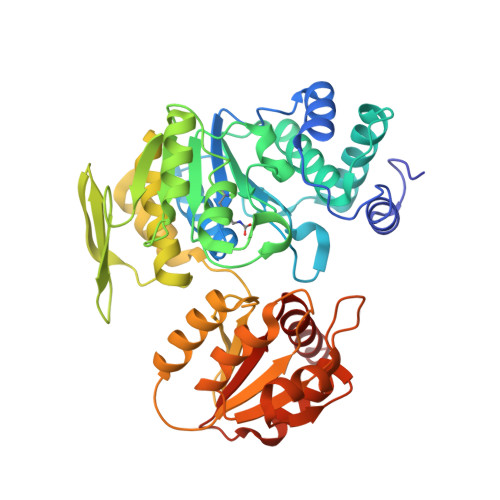
Escherichia Coli Folc Structure Reveals an Unexpected Dihydrofolate Binding Site Providing an Attractive Target for Anti-Microbial Therapy
Mathieu, M., Debousker, G., Vincent, S., Viviani, F., Bamas-Jacques, N., Mikol, V.(2005) J Biological Chem 280: 18916
- PubMed: 15705579 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M413799200
- Primary Citation Related Structures:
1W78, 1W7K - PubMed Abstract:
In some bacteria, such as Escherichia coli, the addition of L-glutamate to dihydropteroate (dihydrofolate synthetase activity) and the subsequent additions of L-glutamate to tetrahydrofolate (folylpolyglutamate synthetase (FPGS) activity) are catalyzed by the same enzyme, FolC. The crystal structure of E. coli FolC is described in this paper. It showed strong similarities to that of the FPGS enzyme of Lactobacillus casei within the ATP binding site and the catalytic site, as do all other members of the Mur synthethase superfamily. FolC structure revealed an unexpected dihydropteroate binding site very different from the folate site identified previously in the FPGS structure. The relevance of this site is exemplified by the presence of phosphorylated dihydropteroate, a reaction intermediate in the DHFS reaction. L. casei FPGS is considered a relevant model for human FPGS. As such, the presence of a folate binding site in E. coli FolC, which is different from the one seen in FPGS enzymes, provides avenues for the design of specific inhibitors of this enzyme in antimicrobial therapy.
- Department of Structural Biology, Aventis Pharma, 13 Quai J. Guesde, F-94403 Vitry/Seine, France. magali.mathieu@sanofi-aventis.com
Organizational Affiliation: