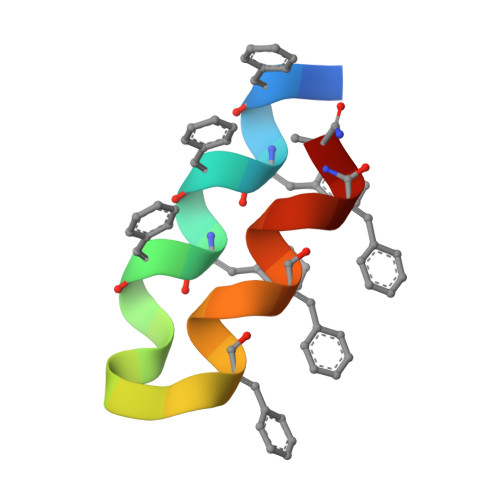
De Novo Design and Characterization of a Helical Hairpin Eicosapeptide; Emergence of an Anion Receptor in the Linker Region.
Rudresh, Ramakumar, S., Ramagopal, U.A., Inai, Y., Goel, S., Sahal, D., Chauhan, V.S.(2004) Structure 12: 389-396
- PubMed: 15016355 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.02.014
- Primary Citation Related Structures:
1VRZ - PubMed Abstract:
De novo design of supersecondary structures is expected to provide useful molecular frameworks for the incorporation of functional sites as in proteins. A 21 residue long, dehydrophenylalanine-containing peptide has been de novo designed and its crystal structure determined. The apolar peptide folds into a helical hairpin supersecondary structure with two right-handed helices, connected by a tetraglycine linker. The helices of the hairpin interact with each other through a combination of C-H.O and N-H.O hydrogen bonds. The folding of the apolar peptide has been realized without the help of either metal ions or disulphide bonds. A remarkable feature of the peptide is the unanticipated occurrence of an anion binding motif in the linker region, strikingly similar in conformation and function to the "nest" motif seen in several proteins. The observation supports the view for the possible emergence of rudimentary functions over short sequence stretches in the early peptides under prebiotic conditions.
- Department of Physics, Indian Institute of Science, Bangalore, India.
Organizational Affiliation: