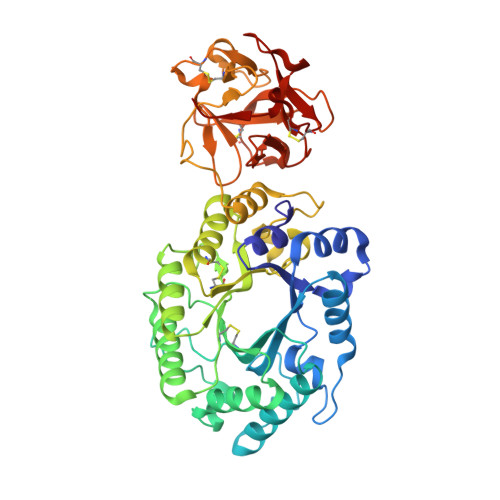
Crystal structures of decorated xylooligosaccharides bound to a family 10 xylanase from Streptomyces olivaceoviridis E-86
Fujimoto, Z., Kaneko, S., Kuno, A., Kobayashi, H., Kusakabe, I., Mizuno, H.(2004) J Biological Chem 279: 9606-9614
- PubMed: 14670957
- DOI: https://doi.org/10.1074/jbc.M312293200
- Primary Citation Related Structures:
1V6U, 1V6V, 1V6W, 1V6X - PubMed Abstract:
The family 10 xylanase from Streptomyces olivaceoviridis E-86 (SoXyn10A) consists of a GH10 catalytic domain, which is joined by a Gly/Pro-rich linker to a family 13 carbohydrate-binding module (CBM13) that interacts with xylan. To understand how GH10 xylanases and CBM13 recognize decorated xylans, the crystal structure of SoXyn10A was determined in complex with alpha-l-arabinofuranosyl- and 4-O-methyl-alpha-d-glucuronosyl-xylooligosaccharides. The bound sugars were observed in the subsites of the catalytic cleft and also in subdomains alpha and gamma of CBM13. The data reveal that the binding mode of the oligosaccharides in the active site of the catalytic domain is entirely consistent with the substrate specificity and, in conjunction with the accompanying paper, demonstrate that the accommodation of the side chains in decorated xylans is conserved in GH10 xylanases of SoXyn10A against arabinoglucuronoxylan. CBM13 was shown to bind xylose or xylooligosaccharides reversibly by using nonsymmetric sugars as the ligands. The independent multiple sites in CBM13 may increase the probability of substrate binding.
- Department of Biochemistry, National Institute of Agrobiological Sciences, Tsukuba, Ibaraki 305-8602, Japan. zui@affrc.go.jp
Organizational Affiliation: