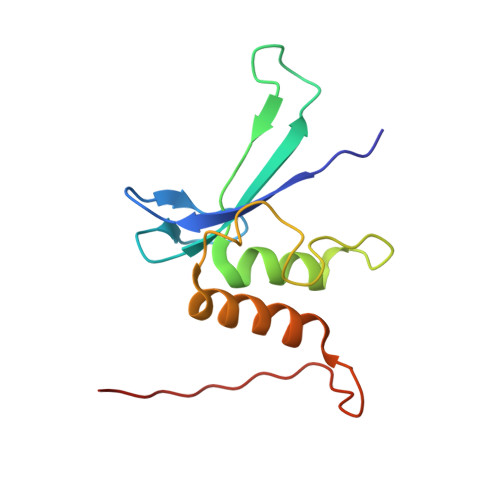
Solution Structure and DNA Binding of the Zinc-Finger Domain from DNA Ligase Iiialpha
Kulczyk, A.W., Yang, J.-C., Neuhaus, D.(2004) J Mol Biology 341: 723
- PubMed: 15288782 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.06.035
- Primary Citation Related Structures:
1UW0 - PubMed Abstract:
DNA ligase IIIalpha carries out the final ligation step in the base excision repair (BER) and single strand break repair (SSBR) mechanisms of DNA repair. The enzyme recognises single-strand nicks and other damage features in double-stranded DNA, both through the catalytic domain and an N-terminal domain containing a single zinc finger. The latter is homologous to other zinc fingers that recognise damaged DNA, two in the N terminus of poly(adenosine-ribose)polymerase and three in the N terminus of the Arabidopsis thaliana nick-sensing DNA 3'-phosphoesterase. Here, we present the solution structure of the zinc-finger domain of human DNA ligase IIIalpha, the first structure of a finger from this group. It is related to that of the erythroid transcription factor GATA-1, but has an additional N-terminal beta-strand and C-terminal alpha-helix. Chemical shift mapping using a DNA ligand containing a single-stranded break showed that the DNA-binding surface of the DNA-ligase IIIalpha zinc finger is substantially different from that of GATA-1, consistent with the fact that the two proteins recognise very different features in the DNA. Likely implications for DNA binding are discussed.
- MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 2QH, UK.
Organizational Affiliation: