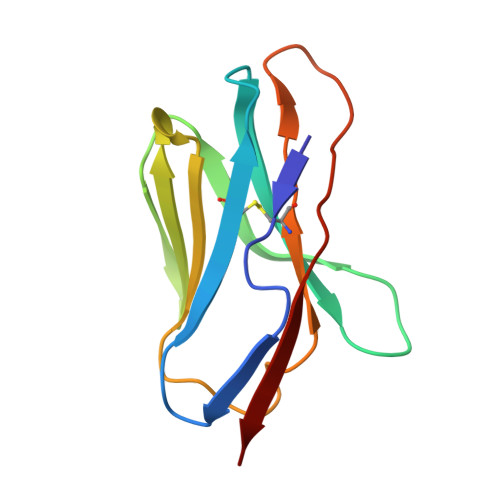
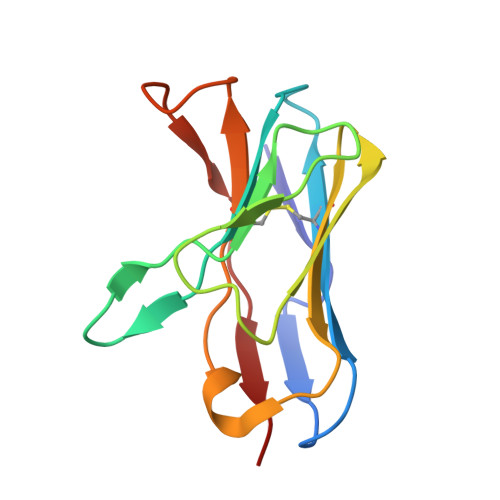
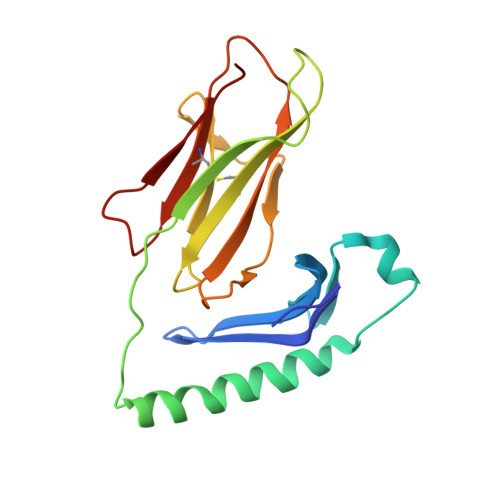
Structure of an autoimmune T cell receptor complexed with class II peptide-MHC: insights into MHC bias and antigen specificity
Maynard, J., Petersson, K., Wilson, D.H., Adams, E.J., Blondelle, S.E., Boulanger, M.J., Wilson, D.B., Garcia, K.C.(2005) Immunity 22: 81-92
- PubMed: 15664161
- DOI: https://doi.org/10.1016/j.immuni.2004.11.015
- Primary Citation of Related Structures:
1U3H - PubMed Abstract:
T cell receptor crossreactivity with different peptide ligands and biased recognition of MHC are coupled features of antigen recognition that are necessary for the T cell's diverse functional repertoire. In the crystal structure between an autoreactive, EAE T cell clone 172.10 and myelin basic protein (1-11) presented by class II MHC I-Au, recognition of the MHC is dominated by the Vbeta domain of the TCR, which interacts with the MHC alpha chain in a manner suggestive of a germline-encoded TCR/MHC "anchor point." Strikingly, there are few specific contacts between the TCR CDR3 loops and the MBP peptide. We also find that over 1,000,000 different peptides derived from combinatorial libraries can activate 172.10, yet the TCR strongly prefers the native MBP contact residues. We suggest that while TCR scanning of pMHC may be degenerate due to the TCR germline bias for MHC, recognition of structurally distinct agonist peptides is not indicative of TCR promiscuity, but rather highly specific alternative solutions to TCR engagement.
- Departments of Microbiology and Immunology, Structural Biology, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: