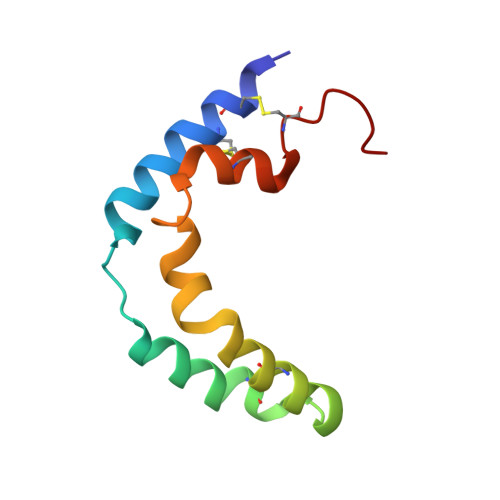
Solution structure of human saposin C in a detergent environment.
Hawkins, C.A., Alba, E., Tjandra, N.(2005) J Mol Biology 346: 1381-1392
- PubMed: 15713488 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.12.045
- Primary Citation Related Structures:
1SN6 - PubMed Abstract:
Saposin C is a lysosomal, membrane-binding protein that acts as an activator for the hydrolysis of glucosylceramide by the enzyme glucocerebrosidase. We used high-resolution NMR to determine the three-dimensional solution structure of saposin C in the presence of the detergent sodium dodecyl sulfate (SDS). This structure provides the first representation of membrane bound saposin C at the atomic level. In the presence of SDS, the protein adopts an open conformation with an exposed hydrophobic pocket. In contrast, the previously reported NMR structure of saposin C in the absence of SDS is compact and contains a hydrophobic core that is not exposed to the solvent. NMR data indicate that the SDS molecules interact with the hydrophobic pocket. The structure of saposin C in the presence of SDS is very similar to a monomer in the saposin B homodimer structure. Their comparison reveals possible similarity in the type of protein/lipid interaction as well as structural components differentiating their quaternary structures and functional specificity.
- Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health, 50 Center Drive, Bethesda, MD 20892, USA.
Organizational Affiliation: