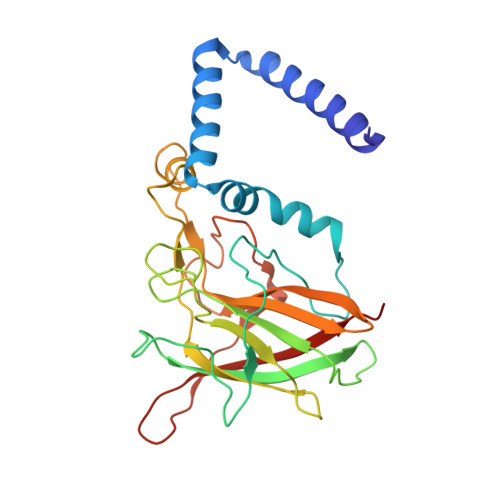
Crystal structure of 4-chlorocatechol 1,2-dioxygenase from the chlorophenol-utilizing gram-positive Rhodococcus opacus 1CP.
Ferraroni, M., Solyanikova, I.P., Kolomytseva, M.P., Scozzafava, A., Golovleva, L., Briganti, F.(2004) J Biological Chem 279: 27646-27655
- PubMed: 15060064 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M401692200
- Primary Citation Related Structures:
1S9A - PubMed Abstract:
The crystal structure of the 4-chlorocatechol 1,2-dioxygenase from the Gram-positive bacterium Rhodococcus opacus (erythropolis) 1CP, a Fe(III) ion-containing enzyme involved in the aerobic biodegradation of chloroaromatic compounds, has been solved by multiple wavelength anomalous dispersion using the weak anomalous signal of the two catalytic irons (1 Fe/257 amino acids) and refined at a 2.5 A resolution (R(free) 28.7%; R factor 21.4%). The analysis of the structure and its comparison with the structure of catechol 1,2-dioxygenase from Acinetobacter calcoaceticus ADP1 (Ac 1,2-CTD) highlight significant differences between these enzymes. The general topology of the present enzyme comprises two catalytic domains (one for each subunit) related by a noncrystallographic 2-fold axis and separated by a common alpha-helical zipper motif consisting of five N-terminal helices from each subunit; furthermore the C-terminal tail is shortened significantly with respect to the known Ac 1,2-CTD. The presence of two phospholipids binding in a hydrophobic tunnel along the dimer axis is shown here to be a common feature for this class of enzyme. The active site cavity presents several dissimilarities with respect to the known catechol-cleaving enzyme. The catalytic nonheme iron(III) ion is bound to the side chains of Tyr-134, Tyr-169, His-194, and His-196, and a cocrystallized benzoate ion, bound to the metal center, reveals details on a novel mode of binding of bidentate inhibitors and a distinctive hydrogen bond network with the surrounding ligands. Among the amino acid residues expected to interact with substrates, several are different from the corresponding analogs of Ac 1,2-CTD: Asp-52, Ala-53, Gly-76, Phe-78, and Cys-224; in addition, regions of largely conserved amino acid residues in the catalytic cleft show different shapes resulting from several substantial backbone and side chain shifts. The present structure is the first of intradiol dioxygenases that specifically catalyze the cleavage of chlorocatechols, key intermediates in the aerobic catabolism of toxic chloroaromatics.
- Dipartimento di Chimica, Università di Firenze, Via della Lastruccia 3, Sesto Fiorentino I-50019, Italy.
Organizational Affiliation: