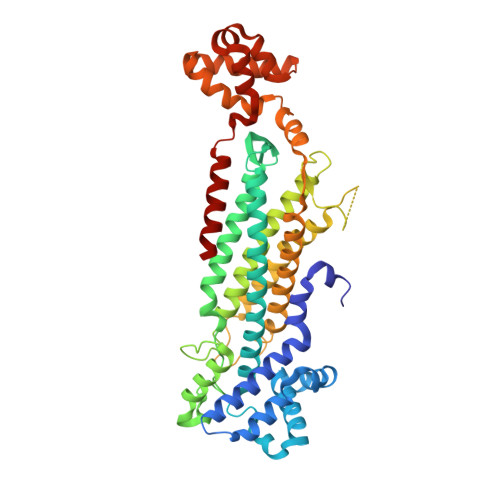
Crystal Structure of 3-Carboxy-cis,cis-muconate Lactonizing Enzyme from Pseudomonas putida, a Fumarase Class II Type Cycloisomerase: Enzyme Evolution in Parallel Pathways.
Yang, J., Wang, Y., Woolridge, E.M., Arora, V., Petsko, G.A., Kozarich, J.W., Ringe, D.(2004) Biochemistry 43: 10424-10434
- PubMed: 15301541
- DOI: https://doi.org/10.1021/bi036205c
- Primary Citation Related Structures:
1RE5 - PubMed Abstract:
3-Carboxy-cis,cis-muconate lactonizing enzymes (CMLEs), the key enzymes in the protocatechuate branch of the beta-ketoadipate pathway in microorganisms, catalyze the conversion of 3-carboxy-cis,cis-muconate to muconolactones. We have determined the crystal structure of the prokaryotic Pseudomonas putida CMLE (PpCMLE) at 2.6 A resolution. PpCMLE is a homotetramer and belongs to the fumarase class II superfamily. The active site of PpCMLE is formed largely by three regions, which are moderately conserved in the fumarase class II superfamily, from three respective monomers. It has been proposed that residue His141, which is highly conserved in all fumarase class II enzymes and forms a charge relay with residue Glu275 (both His141 and Glu275 are in adenylosuccinate lyase numbering), acts as the general base in most fumarase class II superfamily members. However, this charge relay pair is broken in PpCMLE. The residues corresponding to His141 and Glu275 are Trp153 and Ala289, respectively, in PpCMLE. The structures of prokaryotic MLEs and that of CMLE from the eukaryotic Neurospora crassa are completely different from that of PpCMLE, indicating MLEs and CMLEs, as well as the prokaryotic and eukaryotic CMLEs, evolved from distinct ancestors, although they catalyze similar reactions. The structural differences may be related to recognition by substrates and to differences in the mechanistic pathways by which these enzymes catalyze their respective reactions.
- Rosenstiel Basic Medical Sciences Research Center and Department of Biochemistry, Brandeis University, 415 South Street, Waltham, Massachusetts 02454, USA.
Organizational Affiliation: