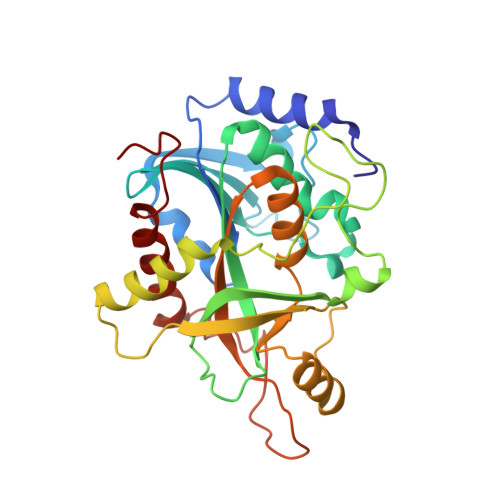
Structures of human purine nucleoside phosphorylase complexed with inosine and ddI
Canduri, F., dos Santos, D.M., Silva, R.G., Mendes, M.A., Basso, L.A., Palma, M.S., de Azevedo Jr., W.F., Santos, D.S.(2004) Biochem Biophys Res Commun 313: 907-914
- PubMed: 14706628 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2003.11.179
- Primary Citation Related Structures:
1RCT, 1V3Q - PubMed Abstract:
Human purine nucleoside phosphorylase (PNP) is a ubiquitous enzyme which plays a key role in the purine salvage pathway, and PNP deficiency in humans leads to an impairment of T-cell function, usually with no apparent effect on B-cell function. PNP is highly specific for 6-oxopurine nucleosides and exhibits negligible activity for 6-aminopurine nucleosides. The catalytic efficiency for inosine is 350,000-fold greater than for adenosine. Adenine nucleosides and nucleotides are deaminated by adenosine deaminase and AMP deaminase to their corresponding inosine derivatives which, in turn, may be further degraded. Here we report the crystal structures of human PNP in complex with inosine and 2('),3(')-dideoxyinosine, refined to 2.8A resolution using synchrotron radiation. The present structures provide explanation for ligand binding, refine the purine-binding site, and can be used for future inhibitor design.
- Departamento de Física, UNESP, São José do Rio Preto, SP 15054-000, Brazil.
Organizational Affiliation: