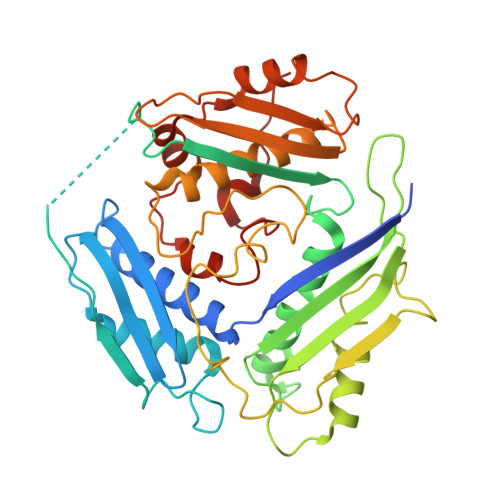
The Crystal Structure of Tetrameric Methionine Adenosyltransferase from Rat Liver Reveals the Methionine-Binding Site
Gonzalez, B., Pajares, M.A., Hermoso, J.A., Alvarez, L., Garrido, F., Sufrin, J.R., Sanz-Aparicio, J.(2000) J Mol Biology 300: 363
- PubMed: 10873471
- DOI: https://doi.org/10.1006/jmbi.2000.3858
- Primary Citation of Related Structures:
1QM4 - PubMed Abstract:
Most of the transmethylation reactions use the same methyl donor, S-adenosylmethionine (SAM), that is synthesised from methionine and ATP by methionine adenosyltransferase (MAT). In mammals, two MAT enzymes have been detected, one ubiquitous and another liver specific. The liver enzyme exists in two oligomeric forms, a tetramer (MAT I) and a dimer (MAT III), MAT I being the one that shows a higher level of affinity for methionine but a lower SAM synthesis capacity. We have solved the crystal structure of rat liver MAT I at 2.7 A resolution, complexed with a methionine analogue: l-2-amino-4-methoxy-cis-but-3-enoic acid (l-cisAMB). The enzyme consists of four identical subunits arranged in two tight dimers that are related by crystallographic 2-fold symmetry. The crystal structure shows the positions of the relevant cysteine residues in the chain, and that Cys35 and Cys61 are perfectly oriented for forming a disulphide link. This result leads us to propose a hypothesis to explain the control of MAT I/III exchange and hence, the effects observed on activity. We have identified the methionine-binding site into the active-site cavity, for the first time. The l-cisAMB inhibitor is stacked against Phe251 aromatic ring in a rather planar conformation, and its carboxylate group coordinates a Mg(2+), which, in turn, is linked to Asp180. The essential role of the involved residues in MAT activity has been confirmed by site-directed mutagenesis. Phe251 is exposed to solvent and is located in the beginning of the flexible loop Phe251-Ala260 that is connecting the N-terminal domain to the central domain. We postulate that a conformational change may take place during the enzymatic reaction and this is possibly the reason of the unusual two-step mechanism involving tripolyphosphate hydrolysis. Other important mechanistic implications are discussed on the light of the results. Moreover, the critical role that certain residues identified in this study may have in methionine recognition opens further possibilities for rational drug design.
- Grupo de Cristalografía Macromolecular y Biología Estructural, Instituto de Química-Física Rocasolano CSIC, Serrano 119, 28006 Madrid, Spain.
Organizational Affiliation: