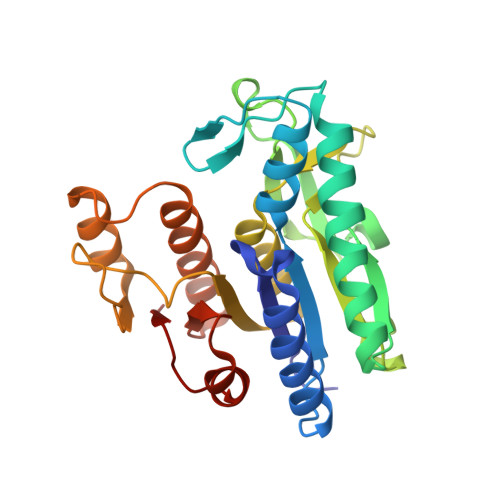
The Course of Phosphorus in the Reaction of N-Acetyl-L-Glutamate Kinase, Determined from the Structures of Crystalline Complexes, Including a Complex with an Alf(4)(-) Transition State Mimic
Gil-Ortiz, F., Ramon-Maiques, S., Fita, I., Rubio, V.(2003) J Mol Biology 331: 231
- PubMed: 12875848 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00716-2
- Primary Citation Related Structures:
1OH9, 1OHA, 1OHB - PubMed Abstract:
N-Acetyl-L-glutamate kinase (NAGK), the structural paradigm of the enzymes of the amino acid kinase family, catalyzes the phosphorylation of the gamma-COO(-) group of N-acetyl-L-glutamate (NAG) by ATP. We determine here the crystal structures of NAGK complexes with MgADP, NAG and the transition-state analog AlF(4)(-); with MgADP and NAG; and with ADP and SO(4)(2-). Comparison of these structures with that of the MgAMPPNP-NAG complex allows to delineate three successive steps during phosphoryl transfer: at the beginning, when the attacking and leaving O atoms and the P atom are imperfectly aligned and the distance between the attacking O atom and the P atom is 2.8A; midway, at the bipyramidal intermediate, with nearly perfect alignment and a distance of 2.3A; and, when the transfer is completed. The transfer occurs in line and is strongly associative, with Lys8 and Lys217 stabilizing the transition state and the leaving group, respectively, and with Lys61, in contrast with an earlier proposal, not being involved. Three water molecules found in all the complexes play, together with Asp162 and the Mg, crucial structural roles. Two glycine-rich loops (beta1-alphaA and beta2-alphaB) are also very important, moving in the different complexes in concert with the ligands, to which they are hydrogen-bonded, either locking them in place for reaction or stabilizing the transition state. The active site is too narrow to accommodate the substrates without compressing the reacting groups, and this compressive strain appears a crucial component of the catalytic mechanism of NAGK, and possibly of other enzymes of the amino acid kinase family such as carbamate kinase. Initial binding of the two substrates would require a different enzyme conformation with a wider active site, and the energy of substrate binding would be used to change the conformation of the active center, causing substrate strain towards the transition state.
- Department of Genomics and Proteomics, Instituto de Biomedicina de Valencia, Consejo Superior de Investigaciones Científicas (IBV-CSIC), C/Jaime Roig 11, 46010- Valencia, Spain.
Organizational Affiliation: