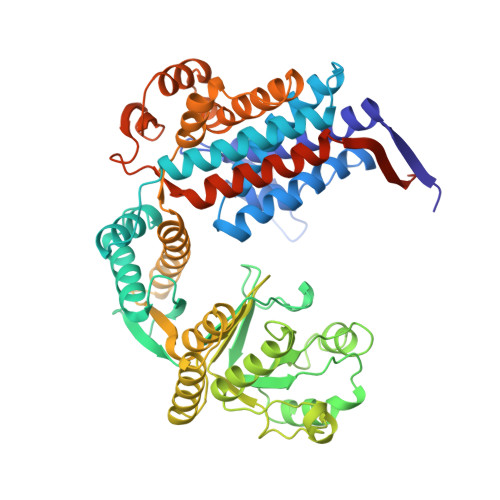
Conformational variability in the refined structure of the chaperonin GroEL at 2.8 A resolution.
Braig, K., Adams, P.D., Brunger, A.T.(1995) Nat Struct Biol 2: 1083-1094
- PubMed: 8846220
- DOI: https://doi.org/10.1038/nsb1295-1083
- Primary Citation Related Structures:
1OEL - PubMed Abstract:
Improved refinement of the crystal structure of GroEL from Escherichia coli has resulted in a complete atomic model for the first 524 residues. A new torsion-angle dynamics method and non-crystallographic symmetry restraints were used in the refinement. The model indicates that conformational variability exists due to rigid-body movements between the apical and intermediate domains of GroEL, resulting in deviations from strict seven-fold symmetry. The regions of the protein involved in polypeptide and GroES binding show unusually high B factors; these values may indicate mobility or discrete disorder. The variability of these regions may play a role in the ability of GroEL to bind a wide variety of substrates.
- Department of Genetics, Yale University, New Haven, Connecticut 06510, USA.
Organizational Affiliation: