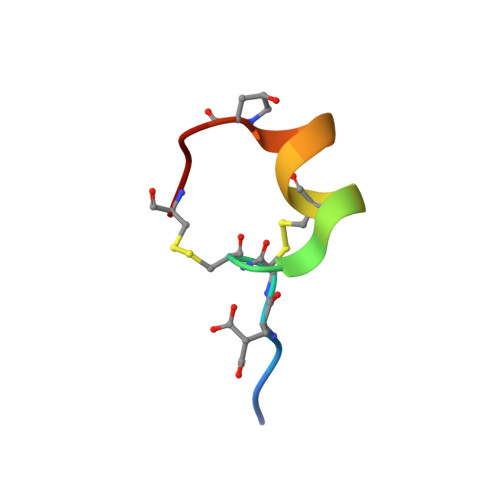
Isolation, Structure, and Activity of GID, a Novel alpha 4/7-Conotoxin with an Extended N-terminal Sequence
Nicke, A., Loughnan, M.L., Millard, E.L., Alewood, P.F., Adams, D.J., Daly, N.L., Craik, D.J., Lewis, R.J.(2003) J Biological Chem 278: 3137-3144
- PubMed: 12419800
- DOI: https://doi.org/10.1074/jbc.M210280200
- Primary Citation of Related Structures:
1MTQ - PubMed Abstract:
Using assay-directed fractionation of Conus geographus crude venom, we isolated alpha-conotoxin GID, which acts selectively at neuronal nicotinic acetylcholine receptors (nAChRs). Unlike other neuronally selective alpha-conotoxins, alpha-GID has a four amino acid N-terminal tail, gamma-carboxyglutamate (Gla), and hydroxyproline (O) residues, and lacks an amidated C terminus. GID inhibits alpha 7 and alpha 3 beta 2 nAChRs with IC(50) values of 5 and 3 nm, respectively and is at least 1000-fold less potent at the alpha 1 beta 1 gamma delta, alpha 3 beta 4, and alpha 4 beta 4 combinations. GID also potently inhibits the alpha 4 beta 2 subtype (IC(50) of 150 nm). Deletion of the N-terminal sequence (GID Delta 1-4) significantly decreased activity at the alpha 4 beta 2 nAChR but hardly affected potency at alpha 3 beta 2 and alpha 7 nAChRs, despite enhancing the off-rates at these receptors. In contrast, Arg(12) contributed to alpha 4 beta 2 and alpha 7 activity but not to alpha 3 beta 2 activity. The three-dimensional structure of GID is well defined over residues 4-19 with a similar motif to other alpha-conotoxins. However, despite its influence on activity, the tail appears to be disordered in solution. Comparison of GID with other alpha 4/7-conotoxins which possess an NN(P/O) motif in loop II, revealed a correlation between increasing length of the aliphatic side-chain in position 10 (equivalent to 13 in GID) and greater alpha 7 versus alpha 3 beta 2 selectivity.
- Institute for Molecular Bioscience and School of Biomedical Sciences, University of Queensland, Brisbane, Queensland 4072, Australia.
Organizational Affiliation: