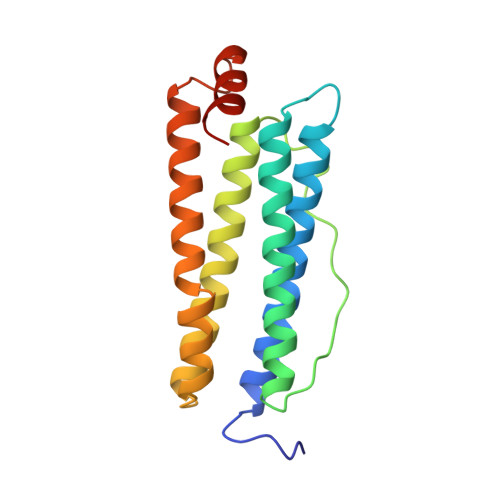
Crystal structure of bullfrog M ferritin at 2.8 A resolution: analysis of subunit interactions and the binuclear metal center
Ha, Y., Shi, D., Small, W., Theil, E.C., Allewell, N.M.(1999) J Biol Inorg Chem 4: 243-256
- PubMed: 10439069 Search on PubMed
- DOI: https://doi.org/10.1007/s007750050310
- Primary Citation Related Structures:
1MFR - PubMed Abstract:
Ferritins concentrate and store iron as a mineral in all bacterial, plant, and animal cells. The two ferritin subunit types, H or M (fast) and L (slow), differ in rates of iron uptake and mineralization and assemble in vivo to form heteropolymeric protein shells made up of 24 subunits; H/L subunit ratios reflect cell specificity of H and L subunit gene expression. A diferric peroxo species that is the initial reaction product of Fe(II) in H-type ferritins, as well as in ribonucleotide reductase (R2) and methane monooxygenase hydroxylase (MMOH), has recently been characterized, exploiting the relatively high accumulation of the peroxo intermediate in frog H-subunit type recombinant ferritin with the M sequence. The stability of the diferric reaction centers in R2 and MMOH contrasts with the instability of diferric centers in ferritin, which are precursors of the ferric mineral. We have determined the crystal structure of the homopolymer of recombinant frog M ferritin in two crystal forms: P4(1)2(1)2, a = b = 170.0 A and c = 481.5 A; and P3(1)21, a = b = 210.8 A and c = 328.1 A. The structural model for the trigonal form was refined to a crystallographic R value of 19.0% (Rfree = 19.4%); the two structures have an r.m.s.d. of approximately 0.22 A for all C alpha atoms. Comparison with the previously determined crystal structure of frog L ferritin indicates that the subunit interface at the molecular twofold axes is most variable, which may relate to the presence of the ferroxidase site in H-type ferritin subunits. Two metal ions (Mg) from the crystallization buffer were found in the ferroxidase site of the M ferritin crystals and interact with Glu23, Glu58, His61, Glu103, Gln137 and, unique to the M subunit, Asp140. The data suggest that Gln137 and Asp140 are a vestige of the second GluxxHis site, resulting from single nucleotide mutations of Glu and His codons and giving rise to Ala140 or Ser140 present in other eukaryotic H-type ferritins, by additional single nucleotide mutations. The observation of the Gln137xxAsp140 site in the frog M ferritin accounts for both the instability of the diferric oxy complexes in ferritin compared to MMOH and R2 and the observed kinetic variability of the diferric peroxo species in different H-type ferritin sequences.
- Department of Biochemistry, University of Minnesota, St. Paul 55108, USA.
Organizational Affiliation: