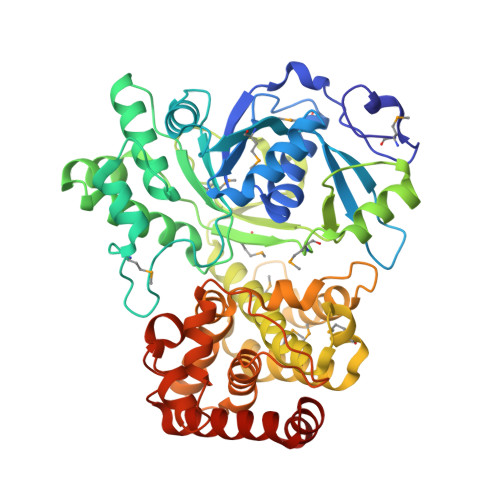
Crystal Structure of Pseudomonas fluorescens Mannitol 2-Dehydrogenase Binary and Ternary Complexes. Specificity and Catalytic Mechanism
Kavanagh, K.L., Klimacek, M., Nidetzky, B., Wilson, D.K.(2002) J Biological Chem 277: 43433-43442
- PubMed: 12196534
- DOI: https://doi.org/10.1074/jbc.M206914200
- Primary Citation of Related Structures:
1LJ8, 1M2W - PubMed Abstract:
Long-chain mannitol dehydrogenases are secondary alcohol dehydrogenases that are of wide interest because of their involvement in metabolism and potential applications in agriculture, medicine, and industry. They differ from other alcohol and polyol dehydrogenases because they do not contain a conserved tyrosine and are not dependent on Zn(2+) or other metal cofactors. The structures of the long-chain mannitol 2-dehydrogenase (54 kDa) from Pseudomonas fluorescens in a binary complex with NAD(+) and ternary complex with NAD(+) and d-mannitol have been determined to resolutions of 1.7 and 1.8 A and R-factors of 0.171 and 0.176, respectively. These results show an N-terminal domain that includes a typical Rossmann fold. The C-terminal domain is primarily alpha-helical and mediates mannitol binding. The electron lone pair of Lys-295 is steered by hydrogen-bonding interactions with the amide oxygen of Asn-300 and the main-chain carbonyl oxygen of Val-229 to act as the general base. Asn-191 and Asn-300 are involved in a web of hydrogen bonding, which precisely orients the mannitol O2 proton for abstraction. These residues also aid in stabilizing a negative charge in the intermediate state and in preventing the formation of nonproductive complexes with the substrate. The catalytic lysine may be returned to its unprotonated state using a rectifying proton tunnel driven by Glu-292 oscillating among different environments. Despite low sequence homology, the closest structural neighbors are glycerol-3-phosphate dehydrogenase, N-(1-d-carboxylethyl)-l-norvaline dehydrogenase, UDP-glucose dehydrogenase, and 6-phosphogluconate dehydrogenase, indicating a possible evolutionary relationship among these enzymes.
- Section of Molecular and Cellular Biology, University of California, Davis, California 95616, USA.
Organizational Affiliation: