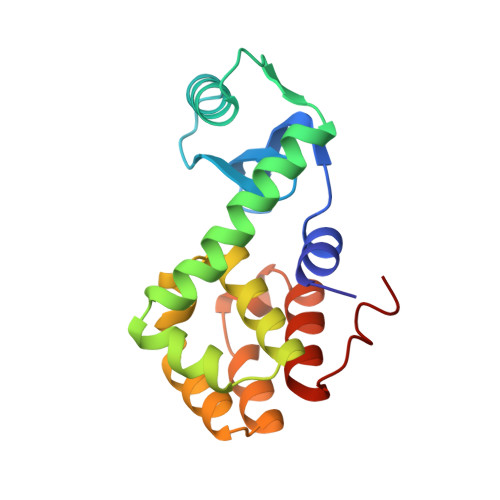
Structural analysis of the temperature-sensitive mutant of bacteriophage T4 lysozyme, glycine 156----aspartic acid.
Gray, T.M., Matthews, B.W.(1987) J Biological Chem 262: 16858-16864
- PubMed: 3680274 Search on PubMed
- DOI: https://doi.org/10.2210/pdb1l16/pdb
- Primary Citation Related Structures:
1L16 - PubMed Abstract:
The structure of the mutant of bacteriophage T4 lysozyme in which Gly-156 is replaced by aspartic acid is described. The lysozyme was isolated by screening for temperature-sensitive mutants and has a melting temperature at pH 6.5 that is 6.1 degrees C lower than wild type. The mutant structure is destabilized, in part, because Gly-156 has conformational angles (phi, psi) that are not optimal for a residue with a beta-carbon. High resolution crystallographic refinement of the mutant structure (R = 17.7% at 1.7 A resolution) shows that the Gly----Asp substitution does not significantly alter the configurational angles (phi, psi) but forces the backbone to move, as a whole, approximately 0.6 A away from its position in wild-type lysozyme. This induced strain weakens a hydrogen bond network that exists in the wild-type structure and also contributes to the reduced stability of the mutant lysozyme. The introduction of an acidic side chain reduces the overall charge on the molecule and thereby tends to increase the stability of the mutant structure relative to wild type. However, at neutral pH this generalized electrostatic stabilization is offset by specific electrostatic repulsion between Asp-156 and Asp-92. The activity of the mutant lysozyme is approximately 50% that of wild-type lysozyme. This reduction in activity might be due to introduction of a negative charge and/or perturbation of the surface of the molecule in the region that is assumed to interact with peptidoglycan substrates.
- Institute of Molecular Biology, University of Oregon, Eugene 97403.
Organizational Affiliation: