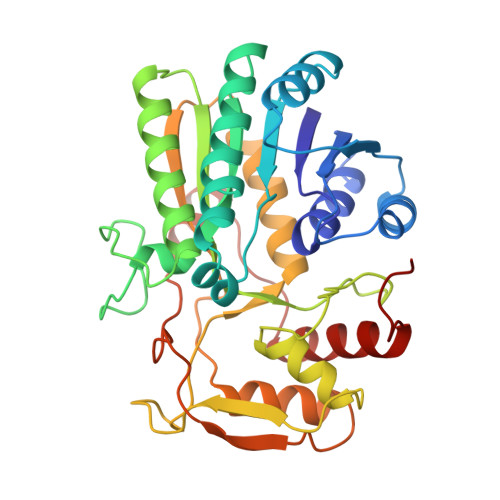
Molecular structures of the S124A, S124T, and S124V site-directed mutants of UDP-galactose 4-epimerase from Escherichia coli.
Thoden, J.B., Gulick, A.M., Holden, H.M.(1997) Biochemistry 36: 10685-10695
- PubMed: 9271499 Search on PubMed
- DOI: https://doi.org/10.1021/bi9704313
- Primary Citation Related Structures:
1KVQ, 1KVR, 1KVS, 1KVT - PubMed Abstract:
UDP-galactose 4-epimerase plays a critical role in sugar metabolism by catalyzing the interconversion of UDP-galactose and UDP-glucose. Originally, it was assumed that the enzyme contained a "traditional" catalytic base that served to abstract a proton from the 4'-hydroxyl group of the UDP-glucose or UDP-galactose substrates during the course of the reaction. However, recent high-resolution X-ray crystallographic analyses of the protein from Escherichia coli have demonstrated the lack of an aspartate, a glutamate, or a histidine residue properly oriented within the active site cleft for serving such a functional role. Rather, the X-ray crystallographic investigation of the epimerase.NADH.UDP-glucose abortive complex from this laboratory has shown that both Ser 124 and Tyr 149 are located within hydrogen bonding distance to the 4'- and 3'-hydroxyl groups of the sugar, respectively. To test the structural role of Ser 124 in the reaction mechanism of epimerase, three site-directed mutant proteins, namely S124A, S124T, and S124V, were constructed and crystals of the S124A.NADH.UDP, S124A.NADH.UDP-glucose, S124T. NADH.UDP-glucose, and S124V.NADH.UDP-glucose complexes were grown. All of the crystals employed in this investigation belonged to the space group P3221 with the following unit cell dimensions: a = b = 83.8 A, c = 108.4 A, and one subunit per asymmetric unit. X-ray data sets were collected to at least 2.15 A resolution, and each protein model was subsequently refined to an R value of lower than 19.0% for all measured X-ray data. The investigations described here demonstrate that the decreases in enzymatic activities observed for these mutant proteins are due to the loss of a properly positioned hydroxyl group at position 124 and not to major tertiary and quaternary structural perturbations. In addition, these structures demonstrate the importance of a hydroxyl group at position 124 in stabilizing the anti conformation of the nicotinamide ring as observed in the previous structural analysis of the epimerase.NADH. UDP complex.
- Institute for Enzyme Research, Graduate School, University of Wisconsin, Madison 53705, USA.
Organizational Affiliation: