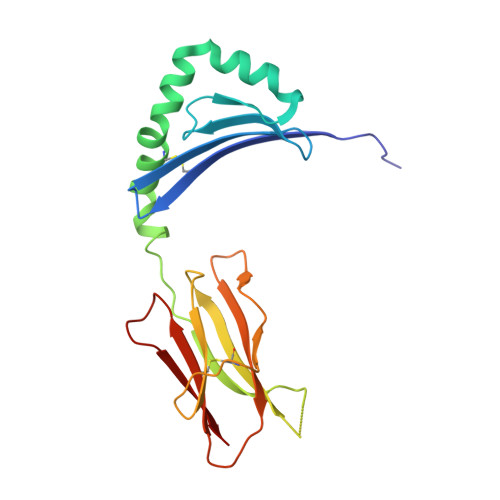
Minor structural changes in a mutated human melanoma antigen correspond to dramatically enhanced stimulation of a CD4+ tumor-infiltrating lymphocyte line.
Sundberg, E.J., Sawicki, M.W., Southwood, S., Andersen, P.S., Sette, A., Mariuzza, R.A.(2002) J Mol Biology 319: 449-461
- PubMed: 12051920
- DOI: https://doi.org/10.1016/S0022-2836(02)00370-4
- Primary Citation Related Structures:
1KLG, 1KLU - PubMed Abstract:
While most immunotherapies for cancer have focused on eliciting specific CD8+ cytotoxic T lymphocyte killing of tumor cells, a mounting body of evidence suggests that stimulation of anti-tumor CD4+ T cell help may be required for highly effective therapy. Several MHC class II-restricted tumor antigens that specifically activate such CD4+ helper T lymphocytes have now been identified, including one from a melanoma tumor that is caused by a single base-pair mutation in the glycolytic enzyme triosephosphate isomerase. This mutation results in the conversion of a threonine residue to isoleucine within the antigenic epitope, concomitant with a greater than five log-fold increase in stimulation of a CD4+ tumor-infiltrating lymphocyte line. Here, we present the crystal structures of HLA-DR1 in complex with both wild-type and mutant TPI peptide antigens, the first structures of tumor peptide antigen/MHC class II complexes recognized by CD4+ T cells to be reported. These structures show that very minor changes in the binding surface for T cell receptor correspond to the dramatic differences in T cell stimulation. Defining the structural basis by which CD4+ T cell help is invoked in an anti-tumor immune response will likely aid the design of more effective cancer immunotherapies.
- Center for Advanced Research in Biotechnology, W. M. Keck Laboratory for Structural Biology, University of Maryland Biotechnology Institute, 9600 Gudelsky Drive, Rockville, MD 20850, USA. sundberg@umbi.umd.edu
Organizational Affiliation: