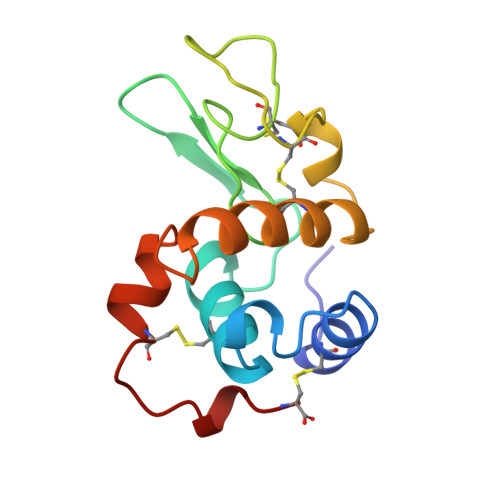
Structure of the calcium-binding echidna milk lysozyme at 1.9 A resolution.
Guss, J.M., Messer, M., Costello, M., Hardy, K., Kumar, V.(1997) Acta Crystallogr D Biol Crystallogr 53: 355-363
- PubMed: 15299900 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444996015831
- Primary Citation Related Structures:
1JUG - PubMed Abstract:
A lysozyme isolated from the milk of a monotreme, the echidna, Tachyglossus aculeatus multiaculeatus, has been crystallized (space group P2(1), with unit-cell dimensions a = 37.1, b = 42.0, c = 38.1 A, beta = 91 degrees and Z = 2) and the structure refined to an R value of 0.167 for all measured data in the resolution range 7.0-1.9 A. It had previously been inferred from sequence homology with alpha-lactalbumins that echidna milk lysozyme (EML) would bind one calcium ion per molecule. This has been confirmed in the present study in which the largest peak in a difference Fourier synthesis is associated with a calcium ion. The calcium binding site of EML is very similar to that observed in baboon and human alpha-lactalbumins, and in a human lysozyme engineered to contain a calcium-binding site. The overall fold of the protein is similar to that of chick-type lysozymes. EML, like pigeon lysozyme, has only 125 residues terminating at a cysteine but in EML this forms a disulfide with a cysteine at residue 9 whereas the equivalent cysteine residue in all other lysozymes of known sequence occurs at position 6. These changes cause some minor structural rearrangements. The binding of calcium appears to have had little effect on the polypeptide backbone conformation and caused only small changes in the conformation of side chains coordinating the calcium ion. A homology modelling study [Acharya, Stuart, Phillips, McKenzie & Teahan (1994). J. Protein Chem. 13(6), 569-584] correctly predicted the overall structure of EML and the nature of its calcium binding site but generally failed to model some more subtle differences observed in the EML structure as evidenced by the fact that the homology model more closely resembles the starting structure from which the model was derived than it does the crystal structure.
- Department of Biochemistry, University of Sydney, NSW, Australia. m.guss@biochem.usyd.edu.au
Organizational Affiliation: