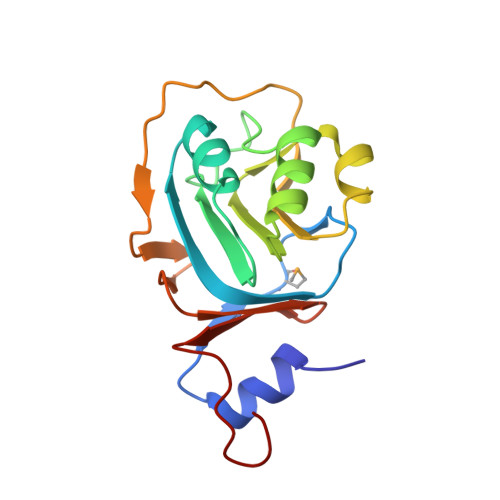
Structure of the RNA-processing inhibitor RraA from Thermus thermophilis.
Rehse, P.H., Kuroishi, C., Tahirov, T.H.(2004) Acta Crystallogr D Biol Crystallogr 60: 1997-2002
- PubMed: 15502308 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904021146
- Primary Citation Related Structures:
1J3L - PubMed Abstract:
The menG gene product, thought to catalyze the final methylation in vitamin K(2) synthesis, has recently been shown to inhibit RNase E in Eschericha coli. The structure of the protein, since renamed RraA, has been solved to 2.3 A using the multiple-wavelength anomalous diffraction method and selenomethionine-substituted protein from Thermus thermophilus. The six molecules in the asymmetric unit are arranged as two similar trimers which have a degree of interaction, suggesting biological significance. The fold does not support the postulated methylation function. Genomic analysis, specifically a lack of an RNase E homologue in cases where homologues to RraA exist, indicates that the function is still obscure.
- Highthroughput Factory, RIKEN Harima Institute, 1-1-1 Kouto, Mikazuki-cho, Sayo-gun, Hyogo 679-5148, Japan.
Organizational Affiliation: