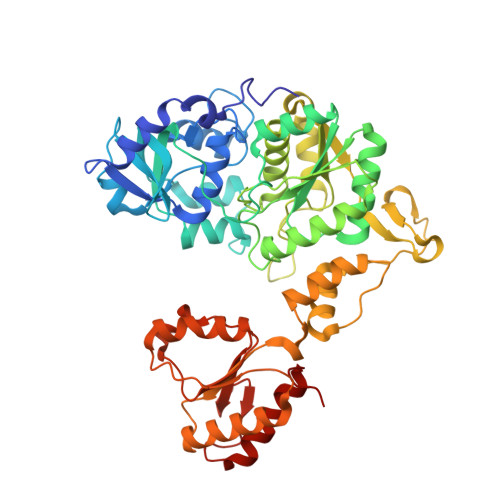
Crystal structure of ATP sulfurylase from Saccharomyces cerevisiae, a key enzyme in sulfate activation.
Ullrich, T.C., Blaesse, M., Huber, R.(2001) EMBO J 20: 316-329
- PubMed: 11157739
- DOI: https://doi.org/10.1093/emboj/20.3.316
- Primary Citation of Related Structures:
1G8F, 1G8G, 1G8H - PubMed Abstract:
ATP sulfurylases (ATPSs) are ubiquitous enzymes that catalyse the primary step of intracellular sulfate activation: the reaction of inorganic sulfate with ATP to form adenosine-5'-phosphosulfate (APS) and pyrophosphate (PPi). With the crystal structure of ATPS from the yeast Saccharomyces cerevisiae, we have solved the first structure of a member of the ATP sulfurylase family. We have analysed the crystal structure of the native enzyme at 1.95 Angstroms resolution using multiple isomorphous replacement (MIR) and, subsequently, the ternary enzyme product complex with APS and PPi bound to the active site. The enzyme consists of six identical subunits arranged in two stacked rings in a D:3 symmetric assembly. Nucleotide binding causes significant conformational changes, which lead to a rigid body structural displacement of domains III and IV of the ATPS monomer. Despite having similar folds and active site design, examination of the active site of ATPS and comparison with known structures of related nucleotidylyl transferases reveal a novel ATP binding mode that is peculiar to ATP sulfurylases.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, D-82152 Martinsried, Germany. tullrich@biochem.mpg.de
Organizational Affiliation: