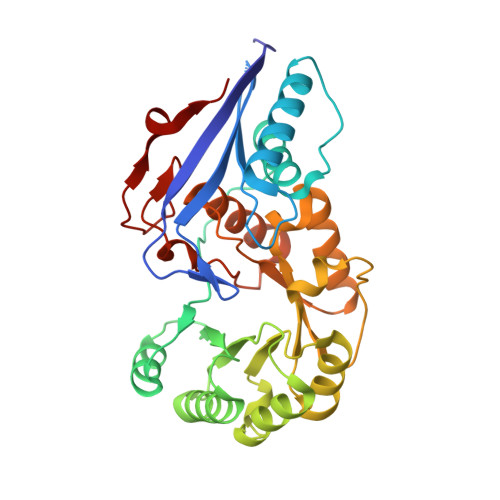
Evolution of enzymatic activity in the enolase superfamily: structure of o-succinylbenzoate synthase from Escherichia coli in complex with Mg2+ and o-succinylbenzoate.
Thompson, T.B., Garrett, J.B., Taylor, E.A., Meganathan, R., Gerlt, J.A., Rayment, I.(2000) Biochemistry 39: 10662-10676
- PubMed: 10978150
- DOI: https://doi.org/10.1021/bi000855o
- Primary Citation Related Structures:
1FHU, 1FHV - PubMed Abstract:
The X-ray structures of the ligand free (apo) and the Mg(2+)*o-succinylbenzoate (OSB) product complex of o-succinylbenzoate synthase (OSBS) from Escherichia coli have been solved to 1.65 and 1.77 A resolution, respectively. The structure of apo OSBS was solved by multiple isomorphous replacement in space group P2(1)2(1)2(1); the structure of the complex with Mg(2+)*OSB was solved by molecular replacement in space group P2(1)2(1)2. The two domain fold found for OSBS is similar to those found for other members of the enolase superfamily: a mixed alpha/beta capping domain formed from segments at the N- and C-termini of the polypeptide and a larger (beta/alpha)(7)beta barrel domain. Two regions of disorder were found in the structure of apo OSBS: (i) the loop between the first two beta-strands in the alpha/beta domain; and (ii) the first sheet-helix pair in the barrel domain. These regions are ordered in the product complex with Mg(2+)*OSB. As expected, the Mg(2+)*OSB pair is bound at the C-terminal end of the barrel domain. The electron density for the phenyl succinate component of the product is well-defined; however, the 1-carboxylate appears to adopt multiple conformations. The metal is octahedrally coordinated by Asp(161), Glu(190), and Asp(213), two water molecules, and one oxygen of the benzoate carboxylate group of OSB. The loop between the first two beta-strands in the alpha/beta motif interacts with the aromatic ring of OSB. Lys(133) and Lys(235) are positioned to function as acid/base catalysts in the dehydration reaction. Few hydrogen bonding or electrostatic interactions are involved in the binding of OSB to the active site; instead, most of the interactions between OSB and the protein are either indirect via water molecules or via hydrophobic interactions. As a result, evolution of both the shape and the volume of the active site should be subject to few structural constraints. This would provide a structural strategy for the evolution of new catalytic activities in homologues of OSBS and a likely explanation for how the OSBS from Amycolaptosis also can catalyze the racemization of N-acylamino acids [Palmer, D. R., Garrett, J. B., Sharma, V., Meganathan, R., Babbitt, P. C., and Gerlt, J. A. (1999) Biochemistry 38, 4252-4258].
- Department of Biochemistry, University of Wisconsin, Madison, Wisconsin 53705, USA.
Organizational Affiliation: