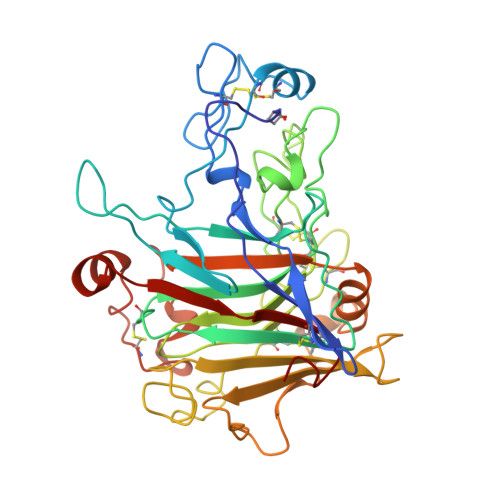
The crystal structure of the catalytic core domain of endoglucanase I from Trichoderma reesei at 3.6 A resolution, and a comparison with related enzymes.
Kleywegt, G.J., Zou, J.Y., Divne, C., Davies, G.J., Sinning, I., Stahlberg, J., Reinikainen, T., Srisodsuk, M., Teeri, T.T., Jones, T.A.(1997) J Mol Biology 272: 383-397
- PubMed: 9325098 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1243
- Primary Citation Related Structures:
1EG1 - PubMed Abstract:
Cellulose is the most abundant polymer in the biosphere. Although generally resistant to degradation, it may be hydrolysed by cellulolytic organisms that have evolved a variety of structurally distinct enzymes, cellobiohydrolases and endoglucanases, for this purpose. Endoglucanase I (EG I) is the major endoglucanase produced by the cellulolytic fungus Trichoderma reesei, accounting for 5 to 10% of the total amount of cellulases produced by this organism. Together with EG I from Humicola insolens and T. reesei cellobiohydrolase I (CBH I), the enzyme is classified into family 7 of the glycosyl hydrolases, and it catalyses hydrolysis with a net retention of the anomeric configuration. The structure of the catalytic core domain (residues 1 to 371) of EG I from T. reesei has been determined at 3.6 A resolution by the molecular replacement method using the structures of T. reesei CBH I and H. insolens EG I as search models. By employing the 2-fold non-crystallographic symmetry (NCS), the structure was refined successfully, despite the limited resolution. The final model has an R-factor of 0.201 (Rfree 0.258). The structure of EG I reveals an extended, open substrate-binding cleft, rather than a tunnel as found in the homologous cellobiohydrolase CBH I. This confirms the earlier proposal that the tunnel-forming loops in CBH I have been deleted in EG I, which has resulted in an open active site in EG I, enabling it to function as an endoglucanase. Comparison of the structure of EG I with several related enzymes reveals structural similarities, and differences that relate to their biological function in degrading particular substrates. A possible structural explanation of the drastically different pH profiles of T. reesei and H. insolens EG I is proposed.
- Biomedical Centre, Uppsala University, Uppsala, SE-751 24, Sweden.
Organizational Affiliation: