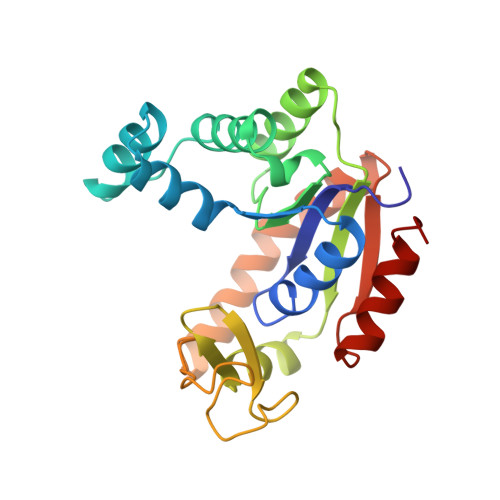
Structure of a mutant adenylate kinase ligated with an ATP-analogue showing domain closure over ATP.
Schlauderer, G.J., Proba, K., Schulz, G.E.(1996) J Mol Biology 256: 223-227
- PubMed: 8594191 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0080
- Primary Citation Related Structures:
1DVR - PubMed Abstract:
Structural studies on unligated and ligated adenylate kinases have shown that two domains, LID and NMPbind, close over the bound substrates, ATP and AMP, respectively. These motions can be, but need not be independent from each other. Up to now, the known structures display only the states "both domains open", "both closed" and "NMP bind closed". In spite of numerous cocrystallization attempts with ATP, a crystalline state "LID closed" has not yet been produced. These experiences suggested that LID closure depends on a bound AMP molecule, in contrast to enzyme kinetic studies indicating a random-bi-bi mechanism. Using an inactive mutant of yeast adenylate kinase together with the ATP analogue AMPPCF2P, however, we have now crystallized an adenylate kinase in the LID closed state. The structure was established at 2.36 A resolution; it indicates that the domain motions occur largely independent from each other in agreement with the kinetic studies. As a side-result, we report the protein environment of the fluorine atoms of the bound ATP analogue.
- Institut für Organische Chemie und Biochemie Albertstr. Freiburg im Breisgau, Germany.
Organizational Affiliation: